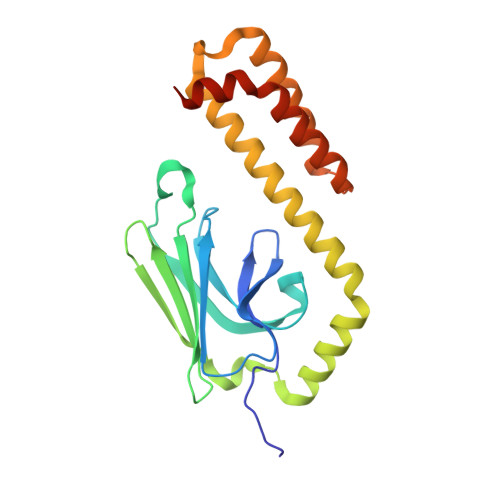
Allosteric opening of the polypeptide-binding site when an Hsp70 binds ATP.
Qi, R., Sarbeng, E.B., Liu, Q., Le, K.Q., Xu, X., Xu, H., Yang, J., Wong, J.L., Vorvis, C., Hendrickson, W.A., Zhou, L., Liu, Q.(2013) Nat Struct Mol Biol 20: 900-907
- PubMed: 23708608 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2583
- Primary Citation Related Structures:
4JN4, 4JNE, 4JNF - PubMed Abstract:
The 70-kilodalton (kDa) heat-shock proteins (Hsp70s) are ubiquitous molecular chaperones essential for cellular protein folding and proteostasis. Each Hsp70 has two functional domains: a nucleotide-binding domain (NBD), which binds and hydrolyzes ATP, and a substrate-binding domain (SBD), which binds extended polypeptides. NBD and SBD interact little when in the presence of ADP; however, ATP binding allosterically couples the polypeptide- and ATP-binding sites. ATP binding promotes polypeptide release; polypeptide rebinding stimulates ATP hydrolysis. This allosteric coupling is poorly understood. Here we present the crystal structure of an intact ATP-bound Hsp70 from Escherichia coli at 1.96-Å resolution. The ATP-bound NBD adopts a unique conformation, forming extensive interfaces with an SBD that has changed radically, having its α-helical lid displaced and the polypeptide-binding channel of its β-subdomain restructured. These conformational changes, together with our biochemical assays, provide a structural explanation for allosteric coupling in Hsp70 activity.
- Department of Physiology and Biophysics, Virginia Commonwealth University, Richmond, Virginia, USA.
Organizational Affiliation: