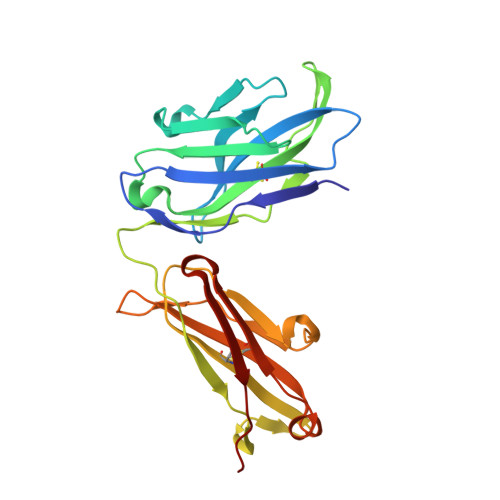
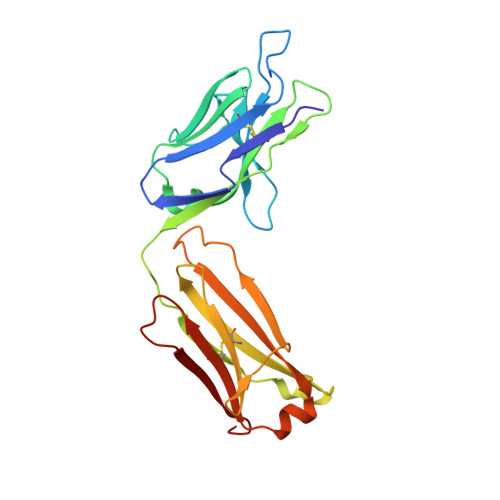
A specific antidote for dabigatran: functional and structural characterization.
Schiele, F., van Ryn, J., Canada, K., Newsome, C., Sepulveda, E., Park, J., Nar, H., Litzenburger, T.(2013) Blood 121: 3554-3562
- PubMed: 23476049 Search on PubMed
- DOI: https://doi.org/10.1182/blood-2012-11-468207
- Primary Citation Related Structures:
4JN1, 4JN2 - PubMed Abstract:
Dabigatran etexilate is a direct thrombin inhibitor and used widely as an anticoagulant for the prevention of stroke in patients with atrial fibrillation. However, anticoagulation therapy can be associated with an increased risk of bleeding. Here, we present data on the identification, humanization, and in vitro pharmacology of an antidote for dabigatran (aDabi-Fab). The X-ray crystal structure of dabigatran in complex with the antidote reveals many structural similarities of dabigatran recognition compared with thrombin. By a tighter network of interactions, the antidote achieves an affinity for dabigatran that is ~350 times stronger than its affinity for thrombin. Despite the structural similarities in the mode of dabigatran binding, the antidote does not bind known thrombin substrates and has no activity in coagulation tests or platelet aggregation. In addition we demonstrate that the antidote rapidly reversed the anticoagulant activity of dabigatran in vivo in a rat model of anticoagulation. This is the first report of a specific antidote for a next-generation anticoagulant that may become a valuable tool in patients who require emergency procedures.
- Structural Research Group, Boehringer Ingelheim GmbH & Co. KG, Biberach, Germany.
Organizational Affiliation: