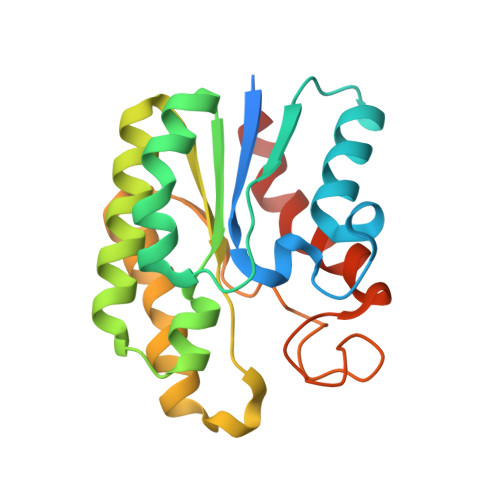
Structural and Functional Characterisation of TesA - A Novel Lysophospholipase A from Pseudomonas aeruginosa.
Kovacic, F., Granzin, J., Wilhelm, S., Kojic-Prodic, B., Batra-Safferling, R., Jaeger, K.E.(2013) PLoS One 8: e69125-e69125
- PubMed: 23874889 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0069125
- Primary Citation Related Structures:
4JGG - PubMed Abstract:
TesA from Pseudomonas aeruginosa belongs to the GDSL hydrolase family of serine esterases and lipases that possess a broad substrate- and regiospecificity. It shows high sequence homology to TAP, a multifunctional enzyme from Escherichia coli exhibiting thioesterase, lysophospholipase A, protease and arylesterase activities. Recently, we demonstrated high arylesterase activity for TesA, but only minor thioesterase and no protease activity. Here, we present a comparative analysis of TesA and TAP at the structural, biochemical and physiological levels. The crystal structure of TesA was determined at 1.9 Å and structural differences were identified, providing a possible explanation for the differences in substrate specificities. The comparison of TesA with other GDSL-hydrolase structures revealed that the flexibility of active-site loops significantly affects their substrate specificity. This assumption was tested using a rational approach: we have engineered the putative coenzyme A thioester binding site of E. coli TAP into TesA of P. aeruginosa by introducing mutations D17S and L162R. This TesA variant showed increased thioesterase activity comparable to that of TAP. TesA is the first lysophospholipase A described for the opportunistic human pathogen P. aeruginosa. The enzyme is localized in the periplasm and may exert important functions in the homeostasis of phospholipids or detoxification of lysophospholipids.
- Institut für Molekulare Enzymtechnologie, Heinrich-Heine Universität Düsseldorf, Forschungszentrum Jülich, Jülich, Germany.
Organizational Affiliation: