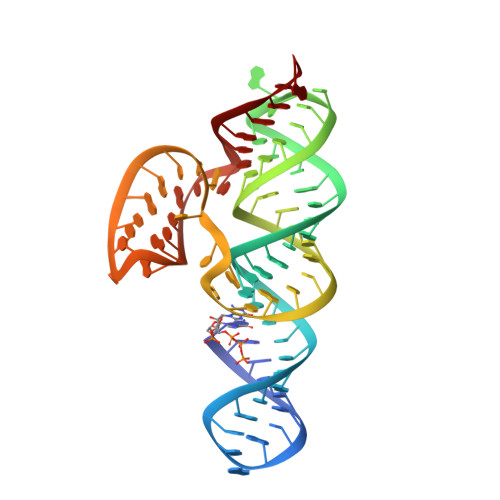
Structure of a class II preQ1 riboswitch reveals ligand recognition by a new fold.
Liberman, J.A., Salim, M., Krucinska, J., Wedekind, J.E.(2013) Nat Chem Biol 9: 353-355
- PubMed: 23584677 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.1231
- Primary Citation Related Structures:
4JF2 - PubMed Abstract:
PreQ1 riboswitches regulate genes by binding the pyrrolopyrimidine intermediate preQ1 during the biosynthesis of the essential tRNA base queuosine. We report what is to our knowledge the first preQ1-II riboswitch structure at 2.3-Å resolution, which uses a previously uncharacterized fold to achieve effector recognition at the confluence of a three-way helical junction flanking a pseudoknotted ribosome-binding site. The results account for translational control mediated by the preQ1-II riboswitch class and expand the known repertoire of ligand-binding modes used by regulatory RNAs.
- Department of Biochemistry and Biophysics, Center for RNA Biology, University of Rochester School of Medicine and Dentistry, Rochester, New York, USA.
Organizational Affiliation: