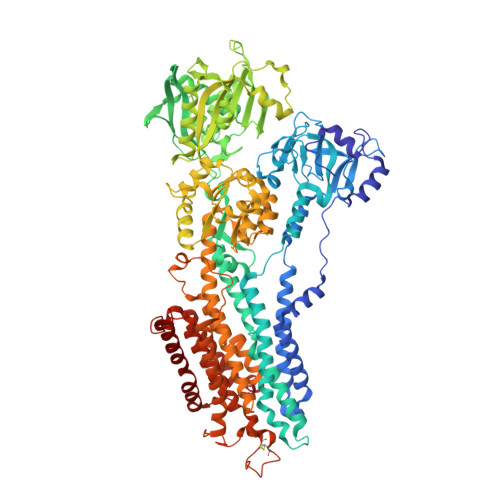
Water-mediated interactions influence the binding of thapsigargin to sarco/endoplasmic reticulum calcium adenosinetriphosphatase.
Paulsen, E.S., Villadsen, J., Tenori, E., Liu, H., Bonde, D.F., Lie, M.A., Bublitz, M., Olesen, C., Autzen, H.E., Dach, I., Sehgal, P., Nissen, P., Moller, J.V., Schiott, B., Christensen, S.B.(2013) J Med Chem 56: 3609-3619
- PubMed: 23574308 Search on PubMed
- DOI: https://doi.org/10.1021/jm4001083
- Primary Citation Related Structures:
4J2T - PubMed Abstract:
A crystal structure suggests four water molecules are present in the binding cavity of thapsigargin in sarco/endoplasmic reticulum calcium ATPase (SERCA). Computational chemistry indicates that three of these water molecules mediate an extensive hydrogen-bonding network between thapsigargin and the backbone of SERCA. The orientation of the thapsigargin molecule in SERCA is crucially dependent on these interactions. The hypothesis has been verified by measuring the affinity of newly synthesized model compounds, which are prevented from participating in such water-mediated interactions as hydrogen-bond donors.
- Department of Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen, Universitetsparken 2, DK-2100 Copenhagen Ø, Denmark.
Organizational Affiliation: