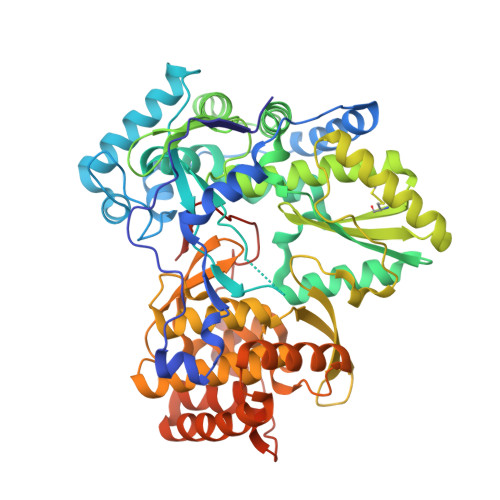
Discovery of a novel series of non-nucleoside thumb pocket 2 HCV NS5B polymerase inhibitors.
Stammers, T.A., Coulombe, R., Rancourt, J., Thavonekham, B., Fazal, G., Goulet, S., Jakalian, A., Wernic, D., Tsantrizos, Y., Poupart, M.A., Bos, M., McKercher, G., Thauvette, L., Kukolj, G., Beaulieu, P.L.(2013) Bioorg Med Chem Lett 23: 2585-2589
- PubMed: 23545108 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2013.02.110
- Primary Citation Related Structures:
4IZ0, 4J02, 4J04, 4J06, 4J08, 4J0A - PubMed Abstract:
A novel series of non-nucleoside thumb pocket 2 HCV NS5B polymerase inhibitors were derived from a fragment-based approach using information from X-ray crystallographic analysis of NS5B-inhibitor complexes and iterative rounds of parallel synthesis. Structure-based drug design strategies led to the discovery of potent sub-micromolar inhibitors 11a-c and 12a-c from a weak-binding fragment-like structure 1 as a starting point.
- Department of Chemistry, Boehringer Ingelheim (Canada) Ltd, Research and Development, 2100 rue Cunard, Laval, Québec, Canada H7S 2G5. timothy.stammers@boehringer-ingelheim.com
Organizational Affiliation: