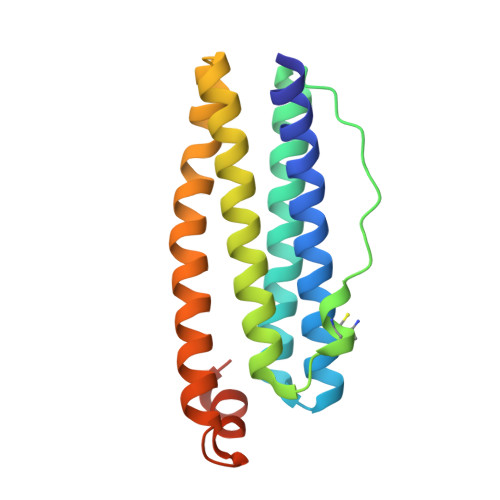
Mechanism of ferrous iron binding and oxidation by ferritin from a pennate diatom.
Pfaffen, S., Abdulqadir, R., Le Brun, N.E., Murphy, M.E.(2013) J Biological Chem 288: 14917-14925
- PubMed: 23548912 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.454496
- Primary Citation Related Structures:
4ISM, 4ISP, 4ITT, 4ITW, 4IWJ, 4IWK, 4IXK - PubMed Abstract:
A novel ferritin was recently found in Pseudo-nitzschia multiseries (PmFTN), a marine pennate diatom that plays a major role in global primary production and carbon sequestration into the deep ocean. Crystals of recombinant PmFTN were soaked in iron and zinc solutions, and the structures were solved to 1.65-2.2-Å resolution. Three distinct iron binding sites were identified as determined from anomalous dispersion data from aerobically grown ferrous soaked crystals. Sites A and B comprise the conserved ferroxidase active site, and site C forms a pathway leading toward the central cavity where iron storage occurs. In contrast, crystal structures derived from anaerobically grown and ferrous soaked crystals revealed only one ferrous iron in the active site occupying site A. In the presence of dioxygen, zinc is observed bound to all three sites. Iron oxidation experiments using stopped-flow absorbance spectroscopy revealed an extremely rapid phase corresponding to Fe(II) oxidation at the ferroxidase site, which is saturated after adding 48 ferrous iron to apo-PmFTN (two ferrous iron per subunit), and a much slower phase due to iron core formation. These results suggest an ordered stepwise binding of ferrous iron and dioxygen to the ferroxidase site in preparation for catalysis and a partial mobilization of iron from the site following oxidation.
- Department of Microbiology and Immunology, University of British Columbia, Vancouver, British Columbia V6T 1Z3, Canada.
Organizational Affiliation: