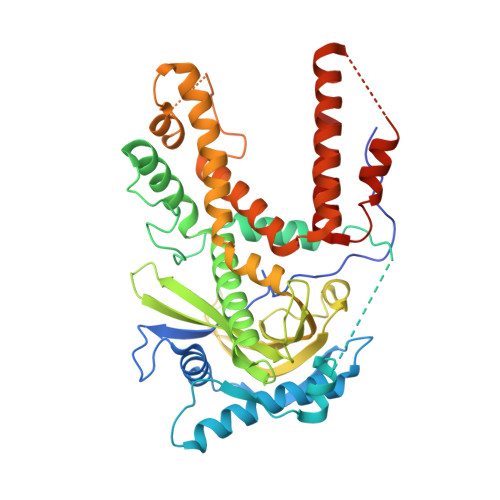
Structural analysis of H1N1 and H7N9 influenza A virus PA in the absence of PB1.
Moen, S.O., Abendroth, J., Fairman, J.W., Baydo, R.O., Bullen, J., Kirkwood, J.L., Barnes, S.R., Raymond, A.C., Begley, D.W., Henkel, G., McCormack, K., Tam, V.C., Phan, I., Staker, B.L., Stacy, R., Myler, P.J., Lorimer, D., Edwards, T.E.(2014) Sci Rep 4: 5944-5944
- PubMed: 25089892 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep05944
- Primary Citation Related Structures:
4IUJ, 4P9A - PubMed Abstract:
Influenza A viruses cause the respiratory illness influenza, which can be mild to fatal depending on the strain and host immune response. The flu polymerase acidic (PA), polymerase basic 1 (PB1), and polymerase basic 2 (PB2) proteins comprise the RNA-dependent RNA polymerase complex responsible for viral genome replication. The first crystal structures of the C-terminal domain of PA (PA-CTD) in the absence of PB1-derived peptides show a number of structural changes relative to the previously reported PB1-peptide bound structures. The human A/WSN/1933 (H1N1) and avian A/Anhui1/2013 (H7N9) strain PA-CTD proteins exhibit the same global topology as other strains in the absence of PB1, but differ extensively in the PB1 binding pocket including a widening of the binding groove and the unfolding of a β-turn. Both PA-CTD proteins exhibited a significant increase in thermal stability in the presence of either a PB1-derived peptide or a previously reported inhibitor in differential scanning fluorimetry assays. These structural changes demonstrate plasticity in the PA-PB1 binding interface which may be exploited in the development of novel therapeutics.
- 1] Seattle Structural Genomics Center for Infectious Disease (SSGCID) [2] Beryllium, 7869 NE Day Road West, Bainbridge Island, WA 98110, USA.
Organizational Affiliation: