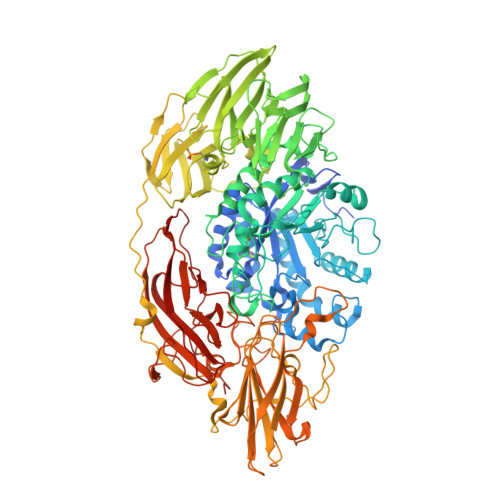
The crystal structure of acidic beta-galactosidase from Aspergillus oryzae.
Maksimainen, M.M., Lampio, A., Mertanen, M., Turunen, O., Rouvinen, J.(2013) Int J Biol Macromol 60C: 109-115
- PubMed: 23688418 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2013.05.003
- Primary Citation Related Structures:
4IUG - PubMed Abstract:
The crystal structure of the industrially important Aspergillus oryzae β-galactosidase has been determined at 2.60 Å resolution. The Ao-β-gal is a large (985 residues) monomeric multi-domain enzyme that has a catalytic (α/β)8-barrel domain. An electron density map revealed extensive N-glycosylation between the domain interfaces suggesting that the oligosaccharide-chains would have a stabilizing role for the structure of Ao-β-gal. Comparison of structure with other β-galactosidase structures of glycoside hydrolase family 35 revealed a number of hydrophobic residues, which may contribute favorably to the stabilization of the structure. The role of a high number of acidic residues in Ao-β-gal is also discussed.
- Department of Chemistry, University of Eastern Finland, P.O. Box 111, 80100 Joensuu, Finland.
Organizational Affiliation: