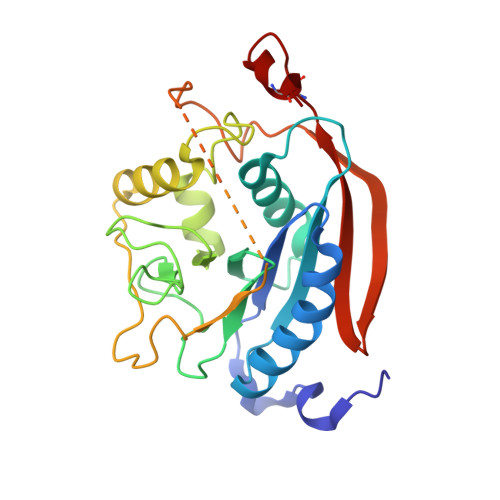
Crystal Structures of beta-1,4-Galactosyltransferase 7 Enzyme Reveal Conformational Changes and Substrate Binding.
Tsutsui, Y., Ramakrishnan, B., Qasba, P.K.(2013) J Biological Chem 288: 31963-31970
- PubMed: 24052259 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.509984
- Primary Citation Related Structures:
4IRP, 4IRQ, 4LW3, 4LW6, 4M4K - PubMed Abstract:
The β-1,4-galactosyltransferase 7 (β4GalT7) enzyme is involved in proteoglycan synthesis. In the presence of a manganese ion, it transfers galactose from UDP-galactose to xylose on a proteoglycan acceptor substrate. We present here the crystal structures of human β4GalT7 in open and closed conformations. A comparison of these crystal structures shows that, upon manganese and UDP or UDP-Gal binding, the enzyme undergoes conformational changes involving a small and a long loop. We also present the crystal structures of Drosophila wild-type β4GalT7 and D211N β4GalT7 mutant enzymes in the closed conformation in the presence of the acceptor substrate xylobiose and the donor substrate UDP-Gal, respectively. To understand the catalytic mechanism, we have crystallized the ternary complex of D211N β4GalT7 mutant enzyme in the presence of manganese with the donor and the acceptor substrates together in the same crystal structure. The galactose moiety of the bound UDP-Gal molecule forms seven hydrogen bonds with the protein molecule. The nonreducing end of the xylose moiety of xylobiose binds to the hydrophobic acceptor sugar binding pocket created by the conformational changes, whereas its extended xylose moiety forms hydrophobic interactions with a Tyr residue. In the ternary complex crystal structure, the nucleophile O4 oxygen atom of the xylose molecule is found in close proximity to the C1 and O5 atoms of the galactose moiety. This is the first time that a Michaelis complex of a glycosyltransferase has been described, and it clearly suggests an SN2 type catalytic mechanism for the β4GalT7 enzyme.
- From the Structural Glycobiology Section and Basic Research Program, SAIC-Frederick, Inc., Nanobiology Program, Center for Cancer Research, NCI, National Institutes of Health, Frederick, Maryland 21702.
Organizational Affiliation: