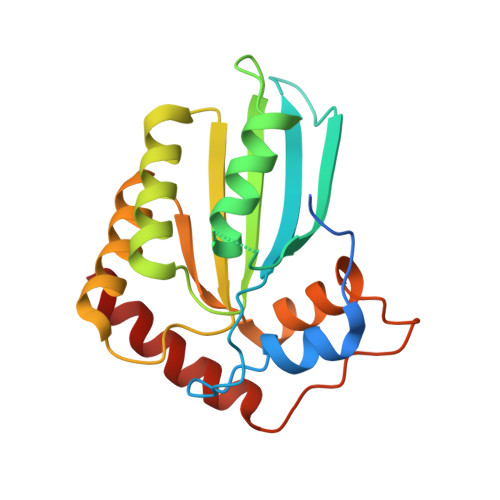
Crystal Structure of the DNA Cytosine Deaminase APOBEC3F: The Catalytically Active and HIV-1 Vif-Binding Domain.
Bohn, M.F., Shandilya, S.M., Albin, J.S., Kouno, T., Anderson, B.D., McDougle, R.M., Carpenter, M.A., Rathore, A., Evans, L., Davis, A.N., Zhang, J., Lu, Y., Somasundaran, M., Matsuo, H., Harris, R.S., Schiffer, C.A.(2013) Structure 21: 1042-1050
- PubMed: 23685212 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2013.04.010
- Primary Citation Related Structures:
4IOU - PubMed Abstract:
Human APOBEC3F is an antiretroviral single-strand DNA cytosine deaminase, susceptible to degradation by the HIV-1 protein Vif. In this study the crystal structure of the HIV Vif binding, catalytically active, C-terminal domain of APOBEC3F (A3F-CTD) was determined. The A3F-CTD shares structural motifs with portions of APOBEC3G-CTD, APOBEC3C, and APOBEC2. Residues identified to be critical for Vif-dependent degradation of APOBEC3F all fit within a predominantly negatively charged contiguous region on the surface of A3F-CTD. Specific sequence motifs, previously shown to play a role in Vif susceptibility and virion encapsidation, are conserved across APOBEC3s and between APOBEC3s and HIV-1 Vif. In this structure these motifs pack against each other at intermolecular interfaces, providing potential insights both into APOBEC3 oligomerization and Vif interactions.
- Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester, MA 01605, USA.
Organizational Affiliation: