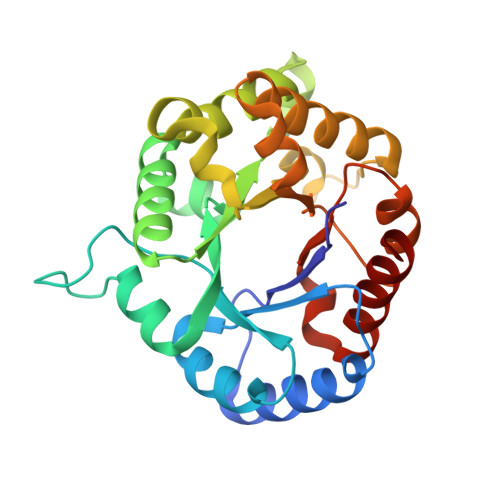
Triosephosphate isomerase is a common crystallization contaminant of soluble His-tagged proteins produced in Escherichia coli.
Kozlov, G., Vinaik, R., Gehring, K.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 499-502
- PubMed: 23695562 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113010841
- Primary Citation Related Structures:
4IOT - PubMed Abstract:
Attempts to crystallize several mammalian proteins overexpressed in Escherichia coli revealed a common contaminant, triosephosphate isomerase, a protein involved in glucose metabolism. Even with triosephosphate isomerase present in very small amounts, similarly shaped crystals appeared in the crystallization drops in a number of polyethylene glycol-containing conditions. All of the target proteins were His-tagged and their purification involved immobilized metal-affinity chromatography (IMAC), a step that was likely to lead to triosephosphate isomerase contamination. Analysis of the triosephosphate isomerase crystals led to the structure of E. coli triosephosphate isomerase at 1.85 Å resolution, which is a significant improvement over the previous structure.
- Department of Biochemistry, Groupe de Recherche axé sur la Structure des Protéines, McGill University, 3649 Promenade Sir William Osler, Montréal, Québec H3G 0B1, Canada.
Organizational Affiliation: