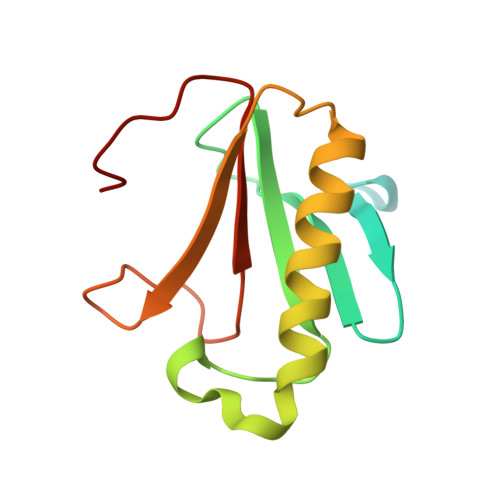
Structural characterization of human histidine triad nucleotide-binding protein 2, a member of the histidine triad superfamily.
Maize, K.M., Wagner, C.R., Finzel, B.C.(2013) FEBS J 280: 3389-3398
- PubMed: 23659632 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/febs.12330
- Primary Citation Related Structures:
4INC, 4INI - PubMed Abstract:
The histidine triad proteins (HITs) constitute a large and ubiquitous superfamily of nucleotide hydrolases. The human histidine triad nucleotide-binding proteins (hHints) are a distinct class of HITs noted for their acyl-AMP hydrolase and phosphoramidase activity. The first high-resolution crystal structures of hHint2 with and without bound AMP are described. The differences between hHint2 and previously known HIT family protein structures are discussed. HIT family enzymes have historically been divided into five classes based on their catalytic specificity: Hint, fragile HIT protein, galactose-1-phosphate uridylyltransferase, DcpS and aprataxin. However, although several structures exist for the enzymes in these classes, the endogenous substrates of many of these enzymes have not been identified or biochemically characterized. To better understand the structural relationships of the HIT enzymes, a structure-based phylogeny was constructed that resulted in the identification of several new putative HIT clades with potential acyl-AMP hydrolase and phosphoramidase activity.
- Department of Medicinal Chemistry, University of Minnesota, Minneapolis, MN, USA.
Organizational Affiliation: