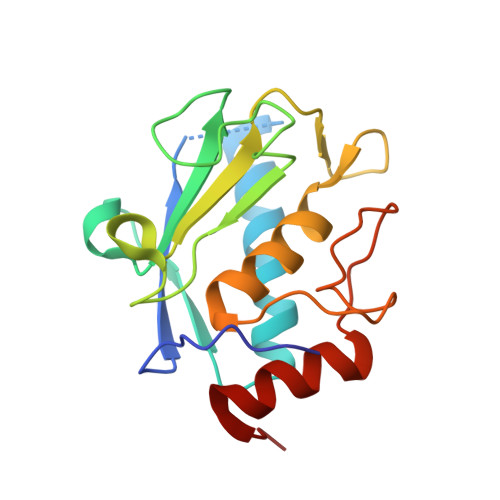
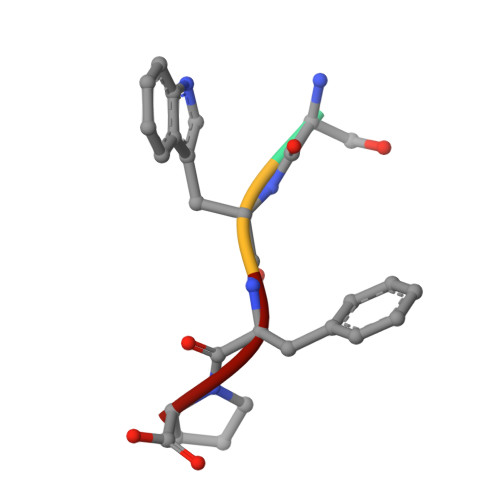
Structure of the catalytic domain of the Tannerella forsythia matrix metallopeptidase karilysin in complex with a tetrapeptidic inhibitor.
Guevara, T., Ksiazek, M., Skottrup, P.D., Cerda-Costa, N., Trillo-Muyo, S., de Diego, I., Riise, E., Potempa, J., Gomis-Ruth, F.X.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 472-476
- PubMed: 23695557 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113007392
- Primary Citation Related Structures:
4IN9 - PubMed Abstract:
Karilysin is the only metallopeptidase identified as a virulence factor in the odontopathogen Tannerella forsythia owing to its deleterious effect on the host immune response during bacterial infection. The very close structural and sequence-based similarity of its catalytic domain (Kly18) to matrix metalloproteinases suggests that karilysin was acquired by horizontal gene transfer from an animal host. Previous studies by phage display identified peptides with the consensus sequence XWFPXXXGGG (single-letter amino-acid codes; X represents any residue) as karilysin inhibitors with low-micromolar binding affinities. Subsequent refinement revealed that inhibition comparable to that of longer peptides could be achieved using the tetrapeptide SWFP. To analyze its binding, the high-resolution crystal structure of the complex between Kly18 and SWFP was determined and it was found that the peptide binds to the primed side of the active-site cleft in a substrate-like manner. The catalytic zinc ion is clamped by the α-amino group and the carbonyl O atom of the serine, thus distantly mimicking the general manner of binding of hydroxamate inhibitors to metallopeptidases and contributing, together with three zinc-binding histidines from the protein scaffold, to an octahedral-minus-one metal-coordination sphere. The tryptophan side chain penetrates the deep partially water-filled specificity pocket of Kly18. Together with previous serendipitous product complexes of Kly18, the present results provide the structural determinants of inhibition of karilysin and open the field for the design of novel inhibitory strategies aimed at the treatment of human periodontal disease based on a peptidic hit molecule.
- Proteolysis Lab, Molecular Biology Institute of Barcelona, Spanish Research Council CSIC, Barcelona Science Park, c/Baldiri Reixac 15-21, 08028 Barcelona, Catalonia, Spain.
Organizational Affiliation: