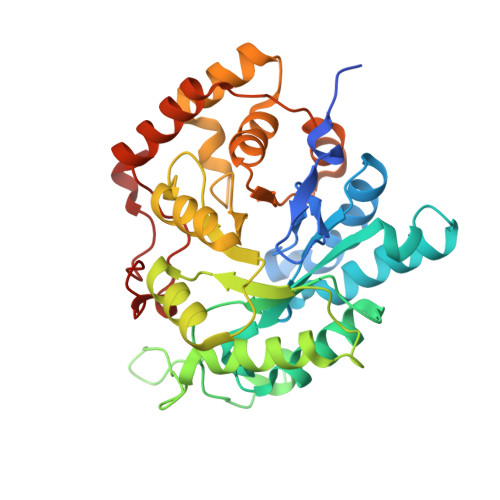
Structures of Saccharomyces cerevisiaeD-arabinose dehydrogenase Ara1 and its complex with NADPH: implications for cofactor-assisted substrate recognition
Hu, X.Q., Guo, P.C., Ma, J.D., Li, W.F.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 1190-1195
- PubMed: 24192347 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113026857
- Primary Citation Related Structures:
4IJC, 4IJR - PubMed Abstract:
The primary role of yeast Ara1, previously mis-annotated as a D-arabinose dehydrogenase, is to catalyze the reduction of a variety of toxic α,β-dicarbonyl compounds using NADPH as a cofactor at physiological pH levels. Here, crystal structures of Ara1 in apo and NADPH-complexed forms are presented at 2.10 and 2.00 Å resolution, respectively. Ara1 exists as a homodimer, each subunit of which adopts an (α/β)8-barrel structure and has a highly conserved cofactor-binding pocket. Structural comparison revealed that induced fit upon NADPH binding yielded an intact active-site pocket that recognizes the substrate. Moreover, the crystal structures combined with computational simulation defined an open substrate-binding site to accommodate various substrates that possess a dicarbonyl group.
- College of Life and Environment Science, Huangshan University, Huangshan, Anhui 245041, People's Republic of China.
Organizational Affiliation: