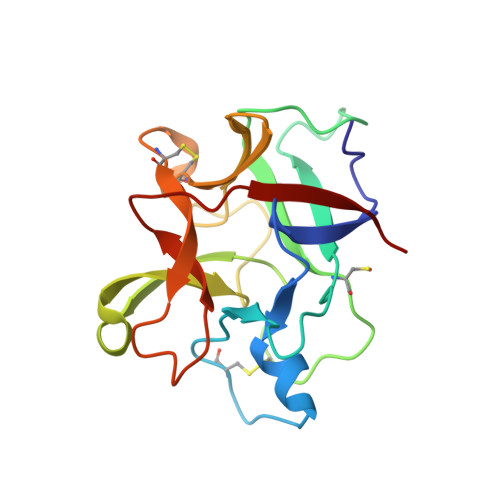
Crystal Structure of Crataeva tapia Bark Protein (CrataBL) and Its Effect in Human Prostate Cancer Cell Lines.
Ferreira, R.D., Zhou, D., Ferreira, J.G., Silva, M.C., Silva-Lucca, R.A., Mentele, R., Paredes-Gamero, E.J., Bertolin, T.C., Dos Santos Correia, M.T., Paiva, P.M., Gustchina, A., Wlodawer, A., Oliva, M.L.(2013) PLoS One 8: e64426-e64426
- PubMed: 23823708 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0064426
- Primary Citation Related Structures:
4IHZ, 4II0 - PubMed Abstract:
A protein isolated from the bark of Crataeva tapia (CrataBL) is both a Kunitz-type plant protease inhibitor and a lectin. We have determined the amino acid sequence and three-dimensional structure of CrataBL, as well as characterized its selected biochemical and biological properties. We found two different isoforms of CrataBL isolated from the original source, differing in positions 31 (Pro/Leu); 92 (Ser/Leu); 93 (Ile/Thr); 95 (Arg/Gly) and 97 (Leu/Ser). CrataBL showed relatively weak inhibitory activity against trypsin (Kiapp = 43 µM) and was more potent against Factor Xa (Kiapp = 8.6 µM), but was not active against a number of other proteases. We have confirmed that CrataBL contains two glycosylation sites and forms a dimer at high concentration. The high-resolution crystal structures of two different crystal forms of isoform II verified the β-trefoil fold of CrataBL and have shown the presence of dimers consisting of two almost identical molecules making extensive contacts (∼645 Å(2)). The structure differs from those of the most closely related proteins by the lack of the N-terminal β-hairpin. In experiments aimed at investigating the biological properties of CrataBL, we have shown that addition of 40 µM of the protein for 48 h caused maximum growth inhibition in MTT assay (47% of DU145 cells and 43% of PC3 cells). The apoptosis of DU145 and PC3 cell lines was confirmed by flow cytometry using Annexin V/FITC and propidium iodide staining. Treatment with CrataBL resulted in the release of mitochondrial cytochrome c and in the activation of caspase-3 in DU145 and PC3 cells.
- Departamento de Bioquímica, Universidade Federal de São Paulo, São Paulo, São Paulo, Brazil.
Organizational Affiliation: