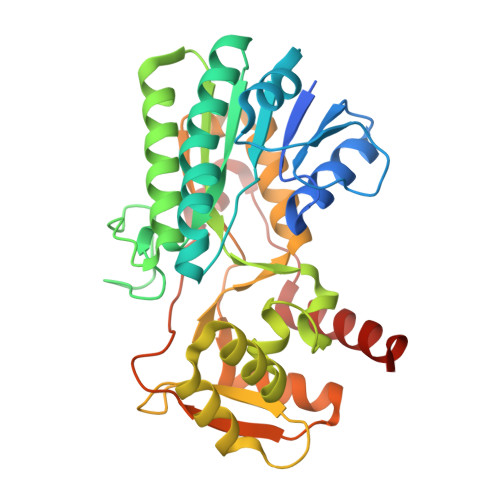
Crystal structure of a short-chain dehydrogenase/reductase superfamily protein from agrobacterium tumefaciens (TARGET EFI-506441) with bound nad, monoclinic form 1
Vetting, M.W., Groninger-Poe, F., Morisco, L.L., Wasserman, S.R., Sojitra, S., Imker, H.J., Gerlt, J.A., Almo, S.C., Enzyme Function Initiative (EFI)To be published.