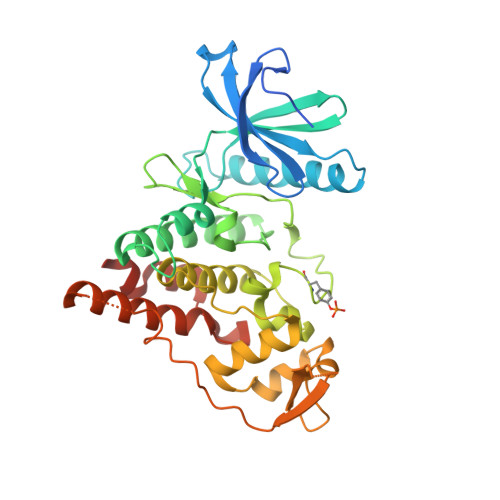
Evaluation of Cancer Dependence and Druggability of PRP4 Kinase Using Cellular, Biochemical, and Structural Approaches.
Gao, Q., Mechin, I., Kothari, N., Guo, Z., Deng, G., Haas, K., McManus, J., Hoffmann, D., Wang, A., Wiederschain, D., Rocnik, J., Czechtizky, W., Chen, X., McLean, L., Arlt, H., Harper, D., Liu, F., Majid, T., Patel, V., Lengauer, C., Garcia-Echeverria, C., Zhang, B., Cheng, H., Dorsch, M., Huang, S.M.(2013) J Biological Chem 288: 30125-30138
- PubMed: 24003220 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.473348
- Primary Citation Related Structures:
4IAN, 4IFC, 4IIR, 4IJP - PubMed Abstract:
PRP4 kinase is known for its roles in regulating pre-mRNA splicing and beyond. Therefore, a wider spectrum of PRP4 kinase substrates could be expected. The role of PRP4 kinase in cancer is also yet to be fully elucidated. Attaining specific and potent PRP4 inhibitors would greatly facilitate the study of PRP4 biological function and its validation as a credible cancer target. In this report, we verified the requirement of enzymatic activity of PRP4 in regulating cancer cell growth and identified an array of potential novel substrates through orthogonal proteomics approaches. The ensuing effort in structural biology unveiled for the first time unique features of PRP4 kinase domain and its potential mode of interaction with a low molecular weight inhibitor. These results provide new and important information for further exploration of PRP4 kinase function in cancer.
- From Discovery and Early Development, Sanofi Oncology, Cambridge, Massachusetts 02139 and 94400 Vitry-sur-Seine Cedex, France.
Organizational Affiliation: