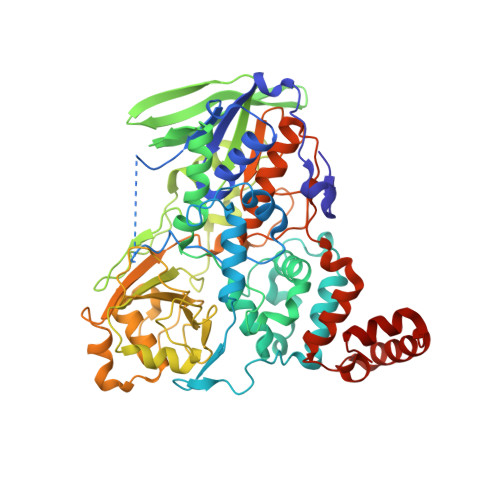
Crystal structure analysis of a fatty acid double-bond hydratase from Lactobacillus acidophilus
Volkov, A., Khoshnevis, S., Neumann, P., Herrfurth, C., Wohlwend, D., Ficner, R., Feussner, I.(2013) Acta Crystallogr D Biol Crystallogr 69: 648-657
- PubMed: 23519674 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913000991
- Primary Citation Related Structures:
4IA5, 4IA6 - PubMed Abstract:
Bacteria have evolved mechanisms for the hydrogenation of unsaturated fatty acids. Hydroxy fatty acid formation may be the first step in such a process; however, knowledge of the structural and mechanistic aspects of this reaction is scarce. Recently, myosin cross-reactive antigen was shown to be a bacterial FAD-containing hydratase which acts on the 9Z and 12Z double bonds of C16 and C18 non-esterified fatty acids, with the formation of 10-hydroxy and 10,13-dihydroxy fatty acids. These fatty acid hydratases form a large protein family which is conserved across Gram-positive and Gram-negative bacteria with no sequence similarity to any known protein apart from the FAD-binding motif. In order to shed light on the substrate recognition and the mechanism of the hydratase reaction, the crystal structure of the hydratase from Lactobacillus acidophilus (LAH) was determined by single-wavelength anomalous dispersion. Crystal structures of apo LAH and of LAH with bound linoleic acid were refined at resolutions of 2.3 and 1.8 Å, respectively. LAH is a homodimer; each protomer consists of four intricately connected domains. Three of them form the FAD-binding and substrate-binding sites and reveal structural similarity to three domains of several flavin-dependent enzymes, including amine oxidoreductases. The additional fourth domain of LAH is located at the C-terminus and consists of three α-helices. It covers the entrance to the hydrophobic substrate channel leading from the protein surface to the active site. In the presence of linoleic acid, the fourth domain of one protomer undergoes conformational changes and opens the entrance to the substrate-binding channel of the other protomer of the LAH homodimer. The linoleic acid molecule is bound at the entrance to the substrate channel, suggesting movement of the lid domain triggered by substrate recognition.
- Department for Plant Biochemistry, Albrecht-von-Haller-Institute for Plant Sciences, Georg-August-University, Göttingen, Germany.
Organizational Affiliation: