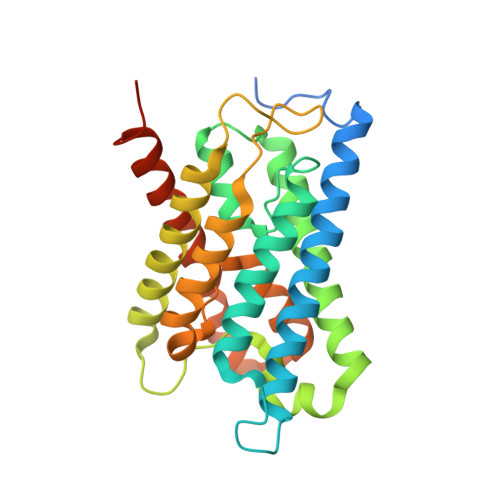
Structural basis for pH gating of plant aquaporins
Frick, A., Jarva, M., Tornroth-Horsefield, S.(2013) FEBS Lett 587: 989-993
- PubMed: 23454640 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2013.02.038
- Primary Citation Related Structures:
4IA4 - PubMed Abstract:
Plants have evolved to cope with fluctuations in water supply by gating their water channels known as aquaporins. During flooding, a rapid drop of cytosolic pH due to anoxia leads to a simultaneous closure of the aquaporins in the plasma membrane. The closing mechanism has been suggested to involve a conserved histidine on cytosolic loop D. Here we report the crystal structure of a spinach aquaporin at low pH, revealing for the first time the structural basis for how this pH-sensitive histidine helps to keep the aquaporin in a closed state.
- Department of Chemistry and Molecular Biology, University of Gothenburg, Gothenburg, Sweden.
Organizational Affiliation: