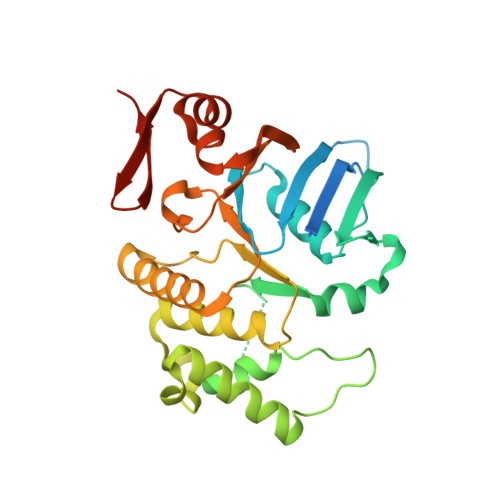
Structural and Functional Characterization of an Orphan ATP-Binding Cassette ATPase Involved in Manganese Utilization and Tolerance in Leptospira spp.
Benaroudj, N., Saul, F., Bellalou, J., Miras, I., Weber, P., Bondet, V., Murray, G.L., Adler, B., Ristow, P., Louvel, H., Haouz, A., Picardeau, M.(2013) J Bacteriol 195: 5583-5591
- PubMed: 24123817 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00915-13
- Primary Citation Related Structures:
4HZI - PubMed Abstract:
Pathogenic Leptospira species are the etiological agents of the widespread zoonotic disease leptospirosis. Most organisms, including Leptospira, require divalent cations for proper growth, but because of their high reactivity, these metals are toxic at high concentrations. Therefore, bacteria have acquired strategies to maintain metal homeostasis, such as metal import and efflux. By screening Leptospira biflexa transposon mutants for their ability to use Mn(2+), we have identified a gene encoding a putative orphan ATP-binding cassette (ABC) ATPase of unknown function. Inactivation of this gene in both L. biflexa and L. interrogans strains led to mutants unable to grow in medium in which iron was replaced by Mn(2+), suggesting an involvement of this ABC ATPase in divalent cation uptake. A mutation in this ATPase-coding gene increased susceptibility to Mn(2+) toxicity. Recombinant ABC ATPase of the pathogen L. interrogans exhibited Mg(2+)-dependent ATPase activity involving a P-loop motif. The structure of this ATPase was solved from a crystal containing two monomers in the asymmetric unit. Each monomer adopted a canonical two-subdomain organization of the ABC ATPase fold with an α/β subdomain containing the Walker motifs and an α subdomain containing the ABC signature motif (LSSGE). The two monomers were arranged in a head-to-tail orientation, forming a V-shaped particle with all the conserved ABC motifs at the dimer interface, similar to functional ABC ATPases. These results provide the first structural and functional characterization of a leptospiral ABC ATPase.
- Institut Pasteur, Unité Biologie des Spirochètes.
Organizational Affiliation: