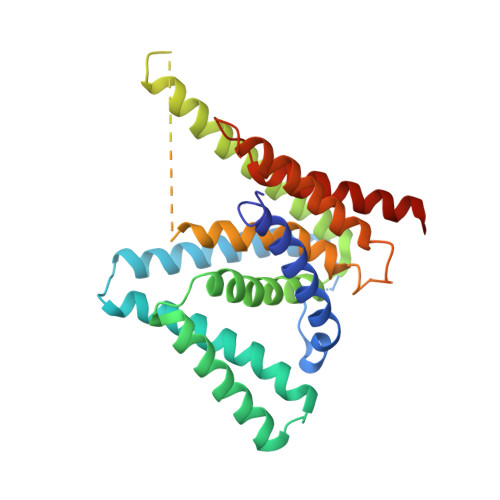
Structure of a presenilin family intramembrane aspartate protease
Li, X., Dang, S., Yan, C., Gong, X., Wang, J., Shi, Y.(2013) Nature 493: 56-61
- PubMed: 23254940 Search on PubMed
- DOI: https://doi.org/10.1038/nature11801
- Primary Citation Related Structures:
4HYC, 4HYD, 4HYG - PubMed Abstract:
Presenilin and signal peptide peptidase (SPP) are intramembrane aspartyl proteases that regulate important biological functions in eukaryotes. Mechanistic understanding of presenilin and SPP has been hampered by lack of relevant structural information. Here we report the crystal structure of a presenilin/SPP homologue (PSH) from the archaeon Methanoculleus marisnigri JR1. The protease, comprising nine transmembrane segments (TMs), adopts a previously unreported protein fold. The amino-terminal domain, consisting of TM1-6, forms a horseshoe-shaped structure, surrounding TM7-9 of the carboxy-terminal domain. The two catalytic aspartate residues are located on the cytoplasmic side of TM6 and TM7, spatially close to each other and approximately 8 Å into the lipid membrane surface. Water molecules gain constant access to the catalytic aspartates through a large cavity between the amino- and carboxy-terminal domains. Structural analysis reveals insights into the presenilin/SPP family of intramembrane proteases.
- Ministry of Education Key Laboratory of Protein Science, Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: