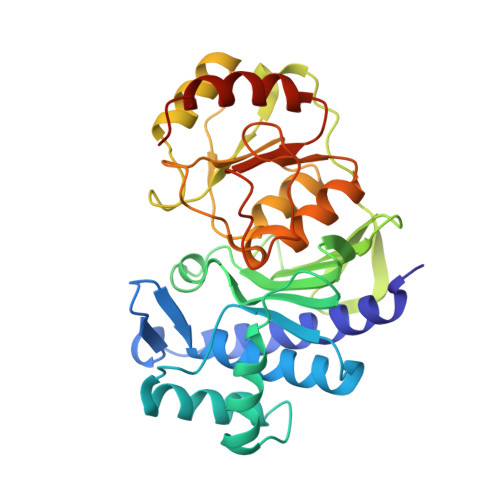
Structural elucidation of a dual-activity PAP phosphatase-1 from Entamoeba histolytica capable of hydrolysing both 3'-phosphoadenosine 5'-phosphate and inositol 1,4-bisphosphate
Faisal Tarique, K., Arif Abdul Rehman, S., Gourinath, S.(2014) Acta Crystallogr D Biol Crystallogr 70: 2019-2031
- PubMed: 25004978 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714010268
- Primary Citation Related Structures:
4HXV, 4O7I - PubMed Abstract:
The enzyme 3'-phosphoadenosine 5'-phosphatase-1 (PAP phosphatase-1) is a member of the Li(+)-sensitive Mg(2+)-dependent phosphatase superfamily, or inositol monophosphatase (IMPase) superfamily, and is an important regulator of the sulfate-activation pathway in all living organisms. Inhibition of this enzyme leads to accumulation of the toxic byproduct 3'-phosphoadenosine 5'-phosphate (PAP), which could be lethal to the organism. Genomic analysis of Entamoeba histolytica suggests the presence of two isoforms of PAP phosphatase. The PAP phosphatase-1 isoform of this organism is shown to be active over wide ranges of pH and temperature. Interestingly, this enzyme is inhibited by submillimolar concentrations of Li(+), while being insensitive to Na(+). Interestingly, the enzyme showed activity towards both PAP and inositol 1,4-bisphosphate and behaved as an inositol polyphosphate 1-phosphatase. Crystal structures of this enzyme in its native form and in complex with adenosine 5'-monophosphate have been determined to 2.1 and 2.6 Å resolution, respectively. The PAP phosphatase-1 structure is divided into two domains, namely α+β and α/β, and the substrate and metal ions bind between them. This is a first structure of any PAP phosphatase to be determined from a human parasitic protozoan. This enzyme appears to function using a mechanism involving three-metal-ion assisted catalysis. Comparison with other structures indicates that the sensitivity to alkali-metal ions may depend on the orientation of a specific catalytic loop.
- School of Life Sciences, Jawaharlal Nehru University, New Delhi 110 067, India.
Organizational Affiliation: