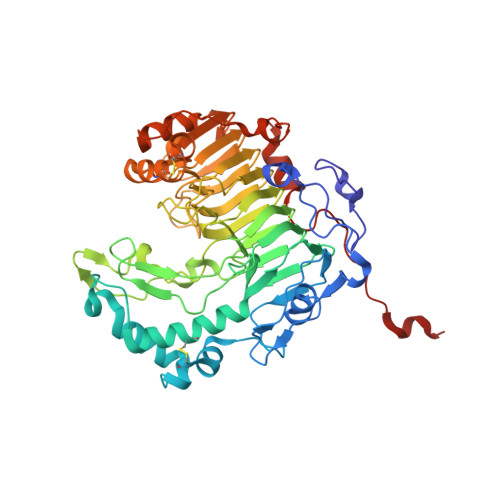
1.37 angstrom crystal structure of pathogenic factor pectate lyase from Acidovorax citrulli.
Tang, Q., Liu, Y.P., Ren, Z.G., Yan, X.X., Zhang, L.Q.(2013) Proteins 81: 1485-1490
- PubMed: 23568384 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24298
- Primary Citation Related Structures:
4HWV - PubMed Abstract:
Pectates lyase (Pel) plays an important role in bacteria pathogenicity. The crystal structure of Pel from Acidovorax citrulli (AcPel) has been solved to 1.37 Å resolution. AcPel belongs to the polysaccharide lyase family 1 (PL1), which has a characteristic right-handed β-helix fold. AcPel is similar with other Pels in the PL1 family, but also shows some differences at the substrate binding site.
- Department of Plant Pathology, China Agricultural University, Beijing, China.
Organizational Affiliation: