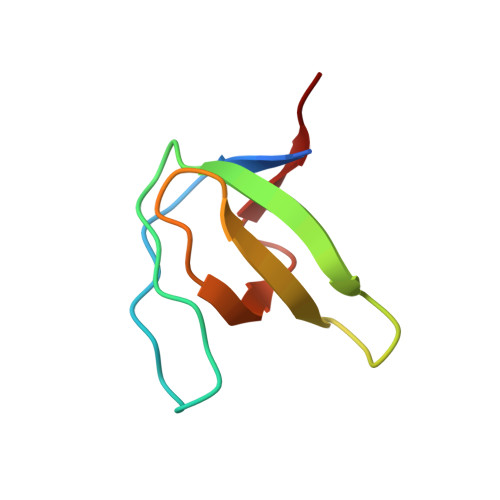
Atomic resolution structures of the c-Src SH3 domain in complex with two high-affinity peptides from classes I and II.
Bacarizo, J., Camara-Artigas, A.(2013) Acta Crystallogr D Biol Crystallogr 69: 756-766
- PubMed: 23633584 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913001522
- Primary Citation Related Structures:
4HVU, 4HVV, 4HVW - PubMed Abstract:
The atomic resolution crystal structures of complexes between the SH3 domain of the c-Src tyrosine kinase and two high-affinity peptides belonging to class I and class II have been solved. The crystals of the Thr98Asp and Thr98Glu mutants in complex with the APP12 peptide (APPLPPRNRPRL) belonged to the trigonal space group P3121 and in both cases the asymmetric unit was composed of one molecule of the SH3-APP12 complex. The crystals of the Thr98Glu mutant in complex with the VSL12 peptide (VSLARRPLPLP) belonged to the trigonal space group P3221 and the asymmetric unit was also composed of a single molecule of the SH3-VSL12 complex. All crystals were obtained in the presence of PEG 300 under the same conditions as reported for the intertwined dimeric structure of the c-Src SH3 domain, but the presence of the peptide stabilizes the monomeric form of the domain. These structures allow a detailed analysis of the role of salt bridges, cation-π interactions and hydrogen bonds in the binding of proline-rich motifs to the c-Src SH3 domain. Moreover, these crystallographic structures allow the role of water molecules in the binding of these motifs to the c-Src SH3 domain to be studied for the first time.
- Department of Chemistry and Physics, University of Almería, Agrifood Campus of International Excellence, Carretera de Sacramento, 04120 Almería, Spain.
Organizational Affiliation: