(4R)- and (4S)-Fluoroproline in the Conserved cis-Prolyl Peptide Bond of the Thioredoxin Fold: Tertiary Structure Context Dictates Ring Puckering.
Rubini, M., Scharer, M.A., Capitani, G., Glockshuber, R.(2013) Chembiochem 14: 1053-1057
- PubMed: 23712956 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.201300178
- Primary Citation Related Structures:
4HU7, 4HU9, 4HUA - PubMed Abstract:
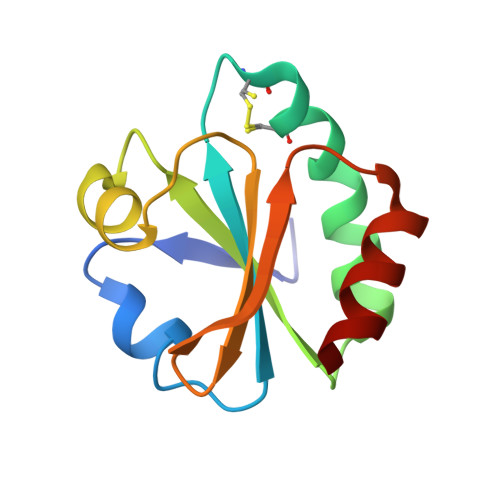
Fine-tuning protein stability: The non-natural amino acids (2S,4R)- and (2S,4S)-fluoroproline modulate protein stability by biasing the proline ring pucker and the cis/trans equilibrium of prolyl peptide bonds. We incorporated both fluoroproline stereoisomers at the invariant cis-proline residue of the thioredoxin fold. The results show that tertiary structure context overrules the conformational preferences of fluoroprolines.
- Department of Organic Chemistry/Cellular Chemistry, University of Konstanz, Universitätsstrasse 10, 78464 Konstanz, Germany. marina.rubini@uni-konstanz.de
Organizational Affiliation: