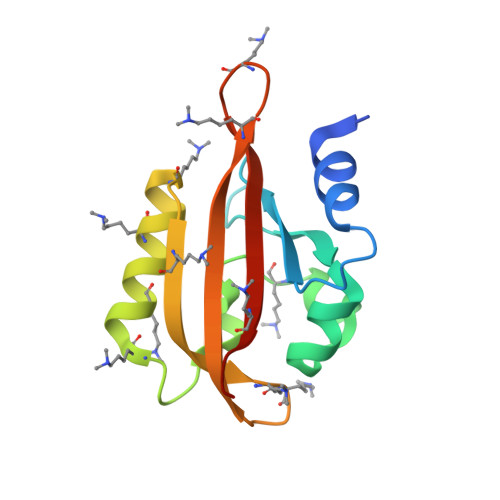
Structural properties of PAS domains from the KCNH potassium channels
Adaixo, R., Harley, C.A., Castro-Rodrigues, A.F., Morais-Cabral, J.H.(2013) PLoS One 8: e59265-e59265
- PubMed: 23555008 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0059265
- Primary Citation Related Structures:
4HOI, 4HP4, 4HP9, 4HQA - PubMed Abstract:
KCNH channels form an important family of voltage gated potassium channels. These channels include a N-terminal Per-Arnt-Sim (PAS) domain with unknown function. In other proteins PAS domains are implicated in cellular responses to environmental queues through small molecule binding or involvement in signaling cascades. To better understand their role we characterized the structural properties of several channel PAS domains. We determined high resolution structures of PAS domains from the mouse EAG (mEAG), drosophila ELK (dELK) and human ERG (hERG) channels and also of the hERG domain without the first nine amino acids. We analyzed these structures for features connected to ligand binding and signaling in other PAS domains. In particular, we have found cavities in the hERG and mEAG structures that share similarities with the ligand binding sites from other PAS domains. These cavities are lined by polar and apolar chemical groups and display potential flexibility in their volume. We have also found that the hydrophobic patch on the domain β-sheet is a conserved feature and appears to drive the formation of protein-protein contacts. In addition, the structures of the dELK domain and of the truncated hERG domain revealed the presence of N-terminal helices. These helices are equivalent to the helix described in the hERG NMR structures and are known to be important for channel function. Overall, these channel domains retain many of the PAS domain characteristics known to be important for cell signaling.
- IBMC, Instituto de Biologia Molecular e Celular, Universidade do Porto, Porto, Portugal.
Organizational Affiliation: