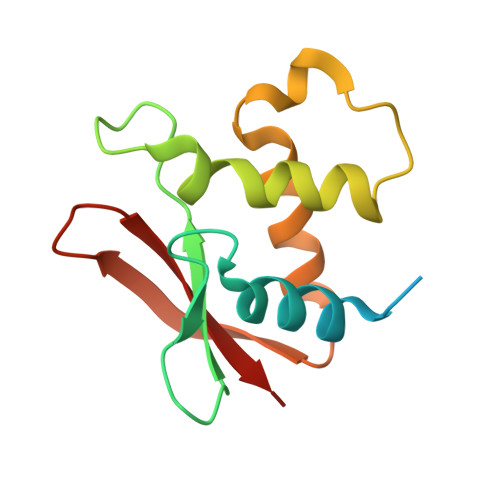
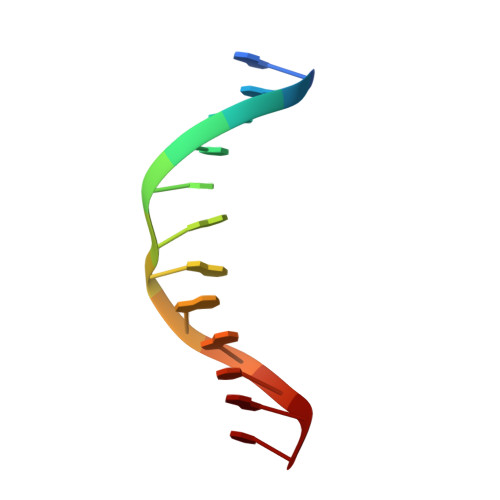
The crystal structure of the DNA-binding domain of vIRF-1 from the oncogenic KSHV reveals a conserved fold for DNA binding and reinforces its role as a transcription factor.
Hew, K., Dahlroth, S.L., Venkatachalam, R., Nasertorabi, F., Lim, B.T., Cornvik, T., Nordlund, P.(2013) Nucleic Acids Res 41: 4295-4306
- PubMed: 23435230 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkt082
- Primary Citation Related Structures:
4HLX, 4HLY - PubMed Abstract:
Kaposi's sarcoma-associated herpesvirus encodes four viral homologues to cellular interferon regulatory factors (IRFs), where the most studied is vIRF-1. Even though vIRF-1 shows sequence homology to the N-terminal DNA-binding domain (DBD) of human IRFs, a specific role for this domain in vIRF-1's function has remained uncertain. To provide insights into the function of the vIRF-1 DBD, we have determined the crystal structure of it in complex with DNA and in its apo-form. Using a thermal stability shift assay (TSSA), we show that the vIRF-1 DBD binds DNA, whereas full-length vIRF-1 does not, suggesting a cis-acting regulatory mechanism in similarity to human IRFs. The complex structure of vIRF-1 DBD reveals interactions with the DNA backbone and the positioning of two arginines for specific recognition in the major grove. A superimposition with human IRF-3 reveals a similar positioning of the two specificity-determining arginines, and additional TSSAs indicate binding of vIRF-1 to an IRF-3 operator consensus sequence. The results from this study, therefore, provide support that vIRF-1 has evolved to bind DNA and plays a role in DNA binding in the context of transcriptional regulation and might act on some of the many operator sequences controlled by human IRF-3.
- Division of Structural Biology and Biochemistry, School of Biological Sciences, Nanyang Technological University, 138673 Singapore, Singapore.
Organizational Affiliation: