Structure reveals function of the dual variable domain immunoglobulin (DVD-Ig[TM]) molecule
Jakob, C.G., Edalji, R., Judge, R.A., Digiammarino, E., Li, Y., Gu, J., Ghayur, T.(2013) MAbs 5: 358-363
- PubMed: 23549062 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4161/mabs.23977
- Primary Citation Related Structures:
4HJJ - PubMed Abstract:
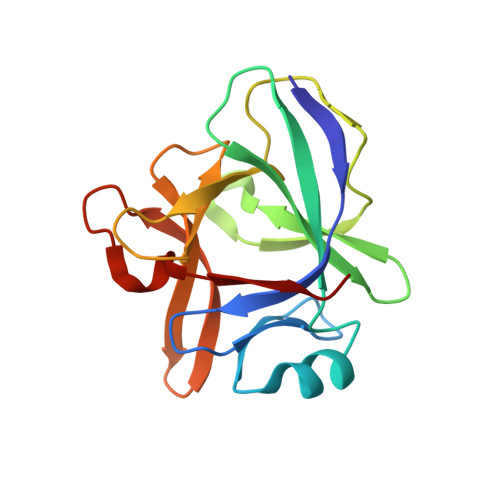
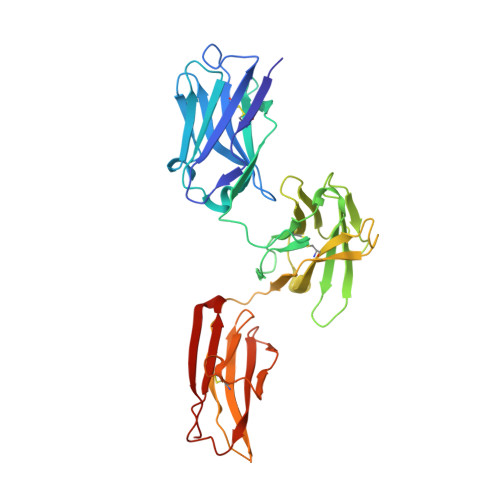
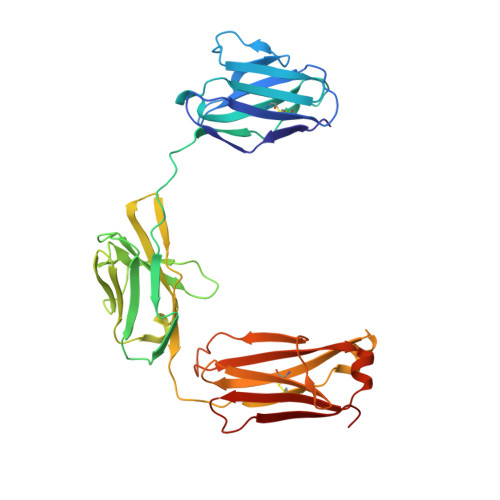
Several bispecific antibody-based formats have been developed over the past 25 years in an effort to produce a new generation of immunotherapeutics that target two or more disease mechanisms simultaneously. One such format, the dual-variable domain immunoglobulin (DVD-Ig™), combines the target binding domains of two monoclonal antibodies via flexible naturally occurring linkers, which yields a tetravalent IgG - like molecule. We report the structure of an interleukin (IL)12-IL18 DVD-Ig™ Fab (DFab) fragment with IL18 bound to the inner variable domain (VD) that reveals the remarkable flexibility of the DVD-Ig™ molecule and how the DVD-Ig™ format can function to bind four antigens simultaneously. An understanding of how the inner variable domain retains function is of critical importance for designing DVD-Ig™ molecules, and for better understanding of the flexibility of immunoglobulin variable domains and linkers, which may aid in the design of improved bi- and multi-specific biologics in general.
- Department of Structural Biology; AbbVie Inc.; North Chicago, IL USA.
Organizational Affiliation: