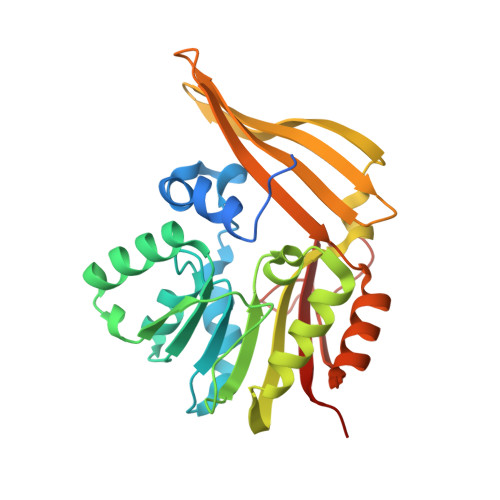
Structure and possible mechanism of the CcbJ methyltransferase from Streptomyces caelestis.
Bauer, J., Ondrovicova, G., Najmanova, L., Pevala, V., Kamenik, Z., Kostan, J., Janata, J., Kutejova, E.(2014) Acta Crystallogr D Biol Crystallogr 70: 943-957
- PubMed: 24699640 Search on PubMed
- DOI: https://doi.org/10.1107/S139900471303397X
- Primary Citation Related Structures:
4HGY, 4HGZ, 4HH4 - PubMed Abstract:
The S-adenosyl-L-methionine (SAM)-dependent methyltransferase CcbJ from Streptomyces caelestis catalyzes one of the final steps in the biosynthesis of the antibiotic celesticetin, methylation of the N atom of its proline moiety, which greatly enhances the activity of the antibiotic. Since several celesticetin variants exist, this enzyme may be able to act on a variety of substrates. The structures of CcbJ determined by MAD phasing at 3.0 Å resolution, its native form at 2.7 Å resolution and its complex with S-adenosyl-L-homocysteine (SAH) at 2.9 Å resolution are reported here. Based on these structures, three point mutants, Y9F, Y17F and F117G, were prepared in order to study its behaviour as well as docking simulations of both CcbJ-SAM-substrate and CcbJ-SAH-product complexes. The structures show that CcbJ is a class I SAM-dependent methyltransferase with a wide active site, thereby suggesting that it may accommodate a number of different substrates. The mutation results show that the Y9F and F117G mutants are almost non-functional, while the Y17F mutant has almost half of the wild-type activity. In combination with the docking studies, these results suggest that Tyr9 and Phe117 are likely to help to position the substrate for the methyl-transfer reaction and that Tyr9 may also facilitate the reaction by removing an H(+) ion. Tyr17, on the other hand, seems to operate by helping to stabilize the SAM cofactor.
- Institute of Molecular Biology, Slovak Academy of Sciences, 851 45 Bratislava, Slovakia.
Organizational Affiliation: