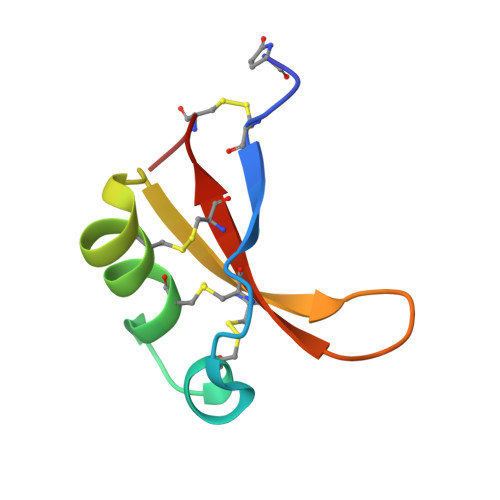
The structure of brazzein, a sweet-tasting protein from the wild African plant Pentadiplandra brazzeana
Nagata, K., Hongo, N., Kameda, Y., Yamamura, A., Sasaki, H., Lee, W.C., Ishikawa, K., Suzuki, E., Tanokura, M.(2013) Acta Crystallogr D Biol Crystallogr 69: 642-647
- PubMed: 23519673 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913001005
- Primary Citation Related Structures:
4HE7 - PubMed Abstract:
Brazzein is the smallest sweet-tasting protein and was isolated from the wild African plant Pentadiplandra brazzeana. The brazzein molecule consists of 54 amino-acid residues and four disulfide bonds. Here, the first crystal structure of brazzein is reported at 1.8 Å resolution and is compared with previously reported solution structures. Despite the overall structural similarity, there are several remarkable differences between the crystal and solution structures both in their backbone folds and side-chain conformations. Firstly, there is an additional α-helix in the crystal structure. Secondly, the atomic r.m.s.d.s between the corresponding C(α)-atom pairs are as large as 2.0-2.2 Å between the crystal and solution structures. Thirdly, the crystal structure exhibits a molecular shape that is similar but not identical to the solution structures. The crystal structure of brazzein reported here will provide additional information and further insights into the intermolecular interaction of brazzein with the sweet-taste receptor.
- Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: