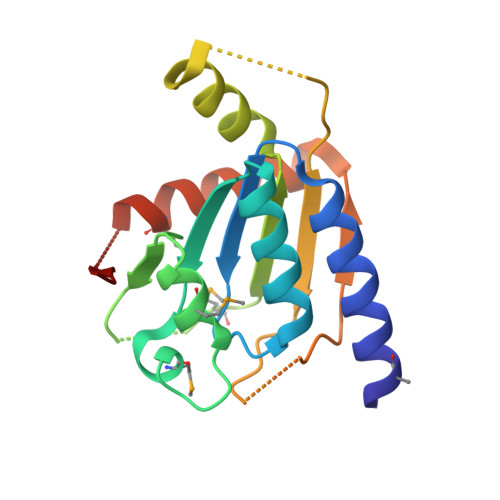
Crystal structure of the primary piRNA biogenesis factor Zucchini reveals similarity to the bacterial PLD endonuclease Nuc.
Voigt, F., Reuter, M., Kasaruho, A., Schulz, E.C., Pillai, R.S., Barabas, O.(2012) RNA 18: 2128-2134
- PubMed: 23086923 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.034967.112
- Primary Citation Related Structures:
4H4A - PubMed Abstract:
Piwi-interacting RNAs (piRNAs) are a gonad-specific class of small RNAs that associate with the Piwi clade of Argonaute proteins and play a key role in transposon silencing in animals. Since biogenesis of piRNAs is independent of the double-stranded RNA-processing enzyme Dicer, an alternative nuclease that can process single-stranded RNA transcripts has been long sought. A Phospholipase D-like protein, Zucchini, that is essential for piRNA processing has been proposed to be a nuclease acting in piRNA biogenesis. Here we describe the crystal structure of Zucchini from Drosophila melanogaster and show that it is very similar to the bacterial endonuclease, Nuc. The structure also reveals that homodimerization induces major conformational changes assembling the active site. The active site is situated on the dimer interface at the bottom of a narrow groove that can likely accommodate single-stranded nucleic acid substrates. Furthermore, biophysical analysis identifies protein segments essential for dimerization and provides insights into regulation of Zucchini's activity.