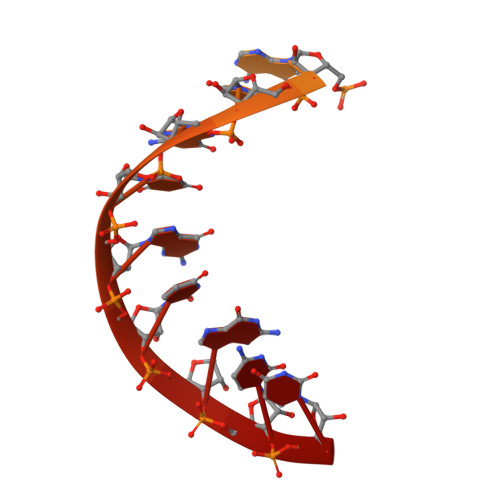
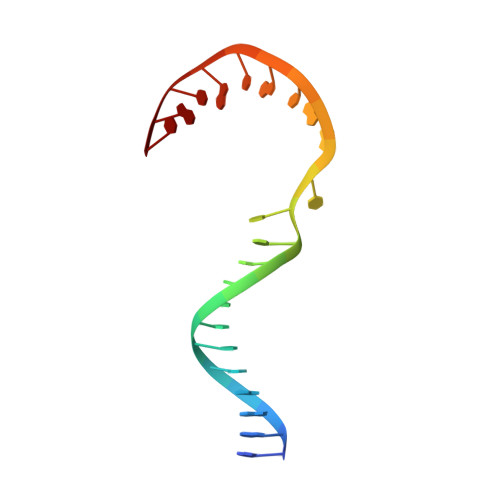
Structural basis of transcriptional pausing in bacteria.
Weixlbaumer, A., Leon, K., Landick, R., Darst, S.A.(2013) Cell 152: 431-441
- PubMed: 23374340 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2012.12.020
- Primary Citation Related Structures:
4GZY, 4GZZ - PubMed Abstract:
Transcriptional pausing by multisubunit RNA polymerases (RNAPs) is a key mechanism for regulating gene expression in both prokaryotes and eukaryotes and is a prerequisite for transcription termination. Pausing and termination states are thought to arise through a common, elemental pause state that is inhibitory for nucleotide addition. We report three crystal structures of Thermus RNAP elemental paused elongation complexes (ePECs). The structures reveal the same relaxed, open-clamp RNAP conformation in the ePEC that may arise by failure to re-establish DNA contacts during translocation. A kinked bridge-helix sterically blocks the RNAP active site, explaining how this conformation inhibits RNAP catalytic activity. Our results provide a framework for understanding how RNA hairpin formation stabilizes the paused state and how the ePEC intermediate facilitates termination.
- The Rockefeller University, 1230 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: