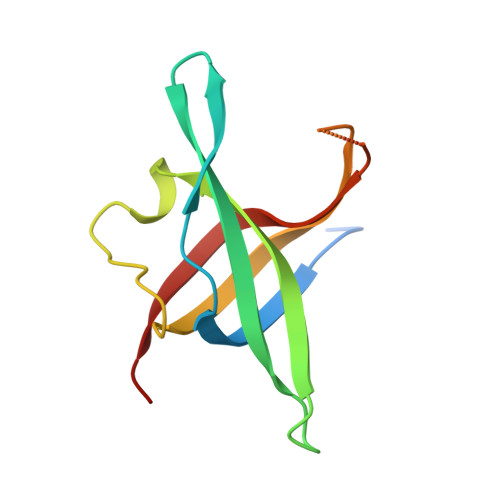
Dimeric structure of the N-terminal domain of PriB protein from Thermoanaerobacter tengcongensis solved ab initio.
Liebschner, D., Brzezinski, K., Dauter, M., Dauter, Z., Nowak, M., Kur, J., Olszewski, M.(2012) Acta Crystallogr D Biol Crystallogr 68: 1680-1689
- PubMed: 23151633 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444912041637
- Primary Citation Related Structures:
4GS3 - PubMed Abstract:
PriB is one of the components of the bacterial primosome, which catalyzes the reactivation of stalled replication forks at sites of DNA damage. The N-terminal domain of the PriB protein from the thermophilic bacterium Thermoanaerobacter tengcongensis (TtePriB) was expressed and its crystal structure was solved at the atomic resolution of 1.09 Å by direct methods. The protein chain, which encompasses the first 104 residues of the full 220-residue protein, adopts the characteristic oligonucleotide/oligosaccharide-binding (OB) structure consisting of a five-stranded β-barrel filled with hydrophobic residues and equipped with four loops extending from the barrel. In the crystal two protomers dimerize, forming a six-stranded antiparallel β-sheet. The structure of the N-terminal OB domain of T. tengcongensis shows significant differences compared with mesophile PriBs. While in all other known structures of PriB a dimer is formed by two identical OB domains in separate chains, TtePriB contains two consecutive OB domains in one chain. However, sequence comparison of both the N-terminal and the C-terminal domains of TtePriB suggests that they have analogous structures and that the natural protein possesses a structure similar to a dimer of two N-terminal domains.
- Synchrotron Radiation Research Section, MCL, National Cancer Institute, Argonne National Laboratory, Argonne, IL 60439, USA.
Organizational Affiliation: