DyP-type peroxidases from Stretptomyces and Thermobifida can modify organosolv lignin.
Lukk, T., Hetta, A.M.A., Jones, A., Solbiati, J., Majumdar, S., Cronan, J.E., Gerlt, J.A., Nair, S.K.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DyP-type peroxidase | 395 | Thermobifida cellulosilytica | Mutation(s): 0 EC: 1.11.1.19 | 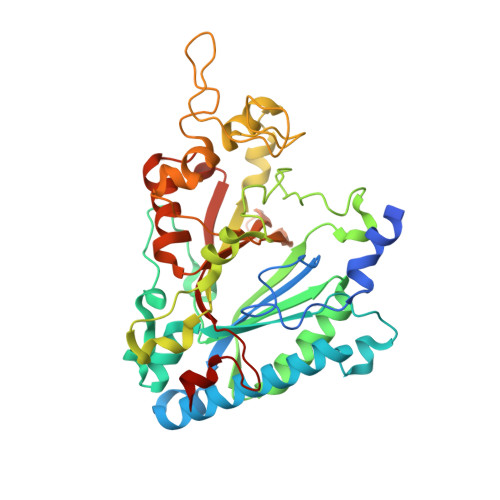 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | U3KRF5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HEM Download:Ideal Coordinates CCD File | C [auth A], F [auth B] | PROTOPORPHYRIN IX CONTAINING FE C34 H32 Fe N4 O4 KABFMIBPWCXCRK-RGGAHWMASA-L |  | ||
| GOL Download:Ideal Coordinates CCD File | E [auth A], H [auth B], I [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| OXY Download:Ideal Coordinates CCD File | D [auth A], G [auth B] | OXYGEN MOLECULE O2 MYMOFIZGZYHOMD-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 87.68 | α = 90 |
| b = 91.19 | β = 90 |
| c = 110.55 | γ = 90 |
| Software Name | Purpose |
|---|---|
| XSCALE | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data collection |
| XDS | data reduction |
| PHASER | phasing |