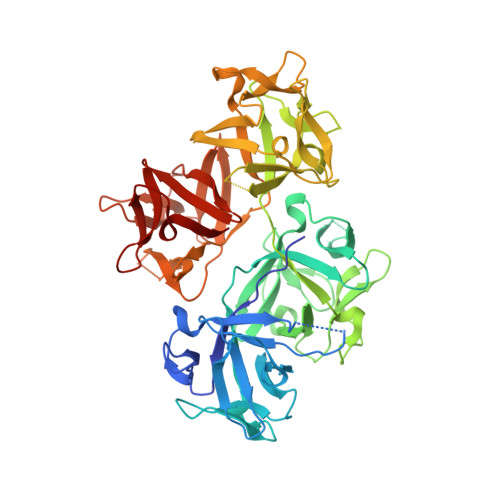
Molecular mechanism of fascin function in filopodial formation.
Yang, S., Huang, F.K., Huang, J., Chen, S., Jakoncic, J., Leo-Macias, A., Diaz-Avalos, R., Chen, L., Zhang, J.J., Huang, X.Y.(2013) J Biological Chem 288: 274-284
- PubMed: 23184945 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.427971
- Primary Citation Related Structures:
4GOV, 4GOY, 4GP0, 4GP3 - PubMed Abstract:
Filopodia are cell surface protrusions that are essential for cell migration. This finger-like structure is supported by rigid tightly bundled actin filaments. The protein responsible for actin bundling in filopodia is fascin. However, the mechanism by which fascin functions in filopodial formation is not clear. Here we provide biochemical, cryo-electron tomographic, and x-ray crystal structural data demonstrating the unique structural characteristics of fascin. Systematic mutagenesis studies on 100 mutants of fascin indicate that there are two major actin-binding sites on fascin. Crystal structures of four fascin mutants reveal concerted conformational changes in fascin from inactive to active states in the process of actin bundling. Mutations in any one of the actin-binding sites impair the cellular function of fascin in filopodial formation. Altogether, our data reveal the molecular mechanism of fascin function in filopodial formation.
- Department of Physiology, Cornell University Weill Medical College, New York, New York 10065, USA.
Organizational Affiliation: