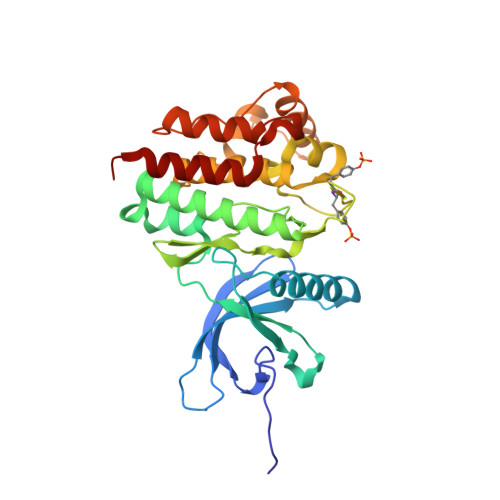
Lead identification of novel and selective TYK2 inhibitors.
Liang, J., Tsui, V., Van Abbema, A., Bao, L., Barrett, K., Beresini, M., Berezhkovskiy, L., Blair, W.S., Chang, C., Driscoll, J., Eigenbrot, C., Ghilardi, N., Gibbons, P., Halladay, J., Johnson, A., Kohli, P.B., Lai, Y., Liimatta, M., Mantik, P., Menghrajani, K., Murray, J., Sambrone, A., Xiao, Y., Shia, S., Shin, Y., Smith, J., Sohn, S., Stanley, M., Ultsch, M., Zhang, B., Wu, L.C., Magnuson, S.(2013) Eur J Med Chem 67: 175-187
- PubMed: 23867602 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejmech.2013.03.070
- Primary Citation Related Structures:
4GFM, 4GFO, 4GIH, 4GMY, 4GVJ - PubMed Abstract:
A therapeutic rationale is proposed for the treatment of inflammatory diseases, such as psoriasis and inflammatory bowel diseases (IBD), by selective targeting of TYK2. Hit triage, following a high-throughput screen for TYK2 inhibitors, revealed pyridine 1 as a promising starting point for lead identification. Initial expansion of 3 separate regions of the molecule led to eventual identification of cyclopropyl amide 46, a potent lead analog with good kinase selectivity, physicochemical properties, and pharmacokinetic profile. Analysis of the binding modes of the series in TYK2 and JAK2 crystal structures revealed key interactions leading to good TYK2 potency and design options for future optimization of selectivity.
- Department of Discovery Chemistry, Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, United States.
Organizational Affiliation: