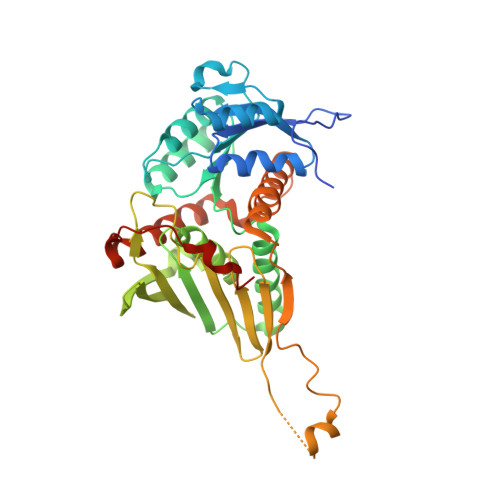
Two Structures of a Thiazolinyl Imine Reductase from Yersinia enterocolitica Provide Insight into Catalysis and Binding to the Nonribosomal Peptide Synthetase Module of HMWP1.
Meneely, K.M., Lamb, A.L.(2012) Biochemistry 51: 9002-9013
- PubMed: 23066849 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi3011016
- Primary Citation Related Structures:
4GMF, 4GMG - PubMed Abstract:
The thiazolinyl imine reductase from Yersinia enterocolitica (Irp3) catalyzes the NADPH-dependent reduction of a thiazoline ring in an intermediate for the formation of the siderophore yersiniabactin. Two structures of Irp3 were determined in the apo (1.85 Å) and NADP(+)-bound (2.31 Å) forms. Irp3 is structurally homologous to sugar oxidoreductases such as glucose-fructose oxidoreductase and 1,5-anhydro-d-fructose reductase, as well as to biliverdin reductase. A homology model of the thiazolinyl imine reductase from Pseudomonas aeruginosa (PchG) was generated. Extensive loop insertions are observed in the C-terminal domain that are unique to Irp3 and PchG and not found in the structural homologues that recognize small molecular substrates. These loops are hypothesized to be important for binding of the nonribosomal peptide synthetase modules (found in HMWP1 and PchF, respectively) to which the substrate of the reductase is covalently attached. A catalytic mechanism for the donation of a proton from a general acid (either histidine 101 or tyrosine 128) and the donation of a hydride from C4 of nicotinamide of the NADPH cofactor is proposed for reduction of the carbon-nitrogen double bond of the thiazoline.
- Department of Molecular Biosciences, University of Kansas, Lawrence, Kansas 66045, United States.
Organizational Affiliation: