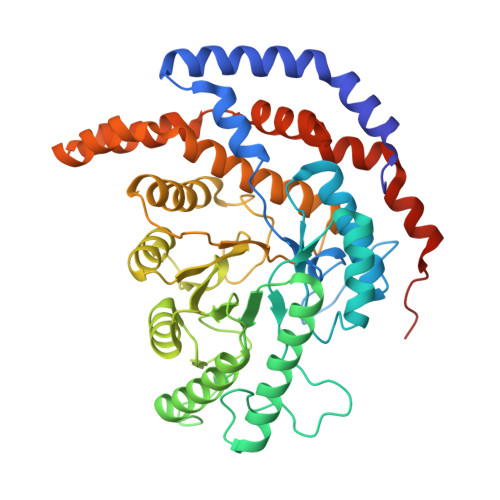
Structure of l-rhamnose isomerase in complex with l-rhamnopyranose demonstrates the sugar-ring opening mechanism and the role of a substrate sub-binding site.
Yoshida, H., Yoshihara, A., Teraoka, M., Yamashita, S., Izumori, K., Kamitori, S.(2013) FEBS Open Bio 3: 35-40
- PubMed: 23772372 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.fob.2012.11.008
- Primary Citation Related Structures:
4GJI, 4GJJ - PubMed Abstract:
l-Rhamnose isomerase (l-RhI) catalyzes the reversible isomerization of l-rhamnose to l-rhamnulose. Previously determined X-ray structures of l-RhI showed a hydride-shift mechanism for the isomerization of substrates in a linear form, but the mechanism for opening of the sugar-ring is still unclear. To elucidate this mechanism, we determined X-ray structures of a mutant l-RhI in complex with l-rhamnopyranose and d-allopyranose. Results suggest that a catalytic water molecule, which acts as an acid/base catalyst in the isomerization reaction, is likely to be involved in pyranose-ring opening, and that a newly found substrate sub-binding site in the vicinity of the catalytic site may recognize different anomers of substrates.
- Life Science Research Center and Faculty of Medicine, Kagawa University, 1750-1 Ikenobe, Miki-cho, Kita-gun, Kagawa 761-0793, Japan.
Organizational Affiliation: