A Molecular Basis for the Control of Preimmune Escape Variants by HIV-Specific CD8(+) T Cells.
Ladell, K., Hashimoto, M., Iglesias, M.C., Wilmann, P.G., McLaren, J.E., Gras, S., Chikata, T., Kuse, N., Fastenackels, S., Gostick, E., Bridgeman, J.S., Venturi, V., Arkoub, Z.A., Agut, H., van Bockel, D.J., Almeida, J.R., Douek, D.C., Meyer, L., Venet, A., Takiguchi, M., Rossjohn, J., Price, D.A., Appay, V.(2013) Immunity 38: 425-436
- PubMed: 23521884 Search on PubMed
- DOI: https://doi.org/10.1016/j.immuni.2012.11.021
- Primary Citation Related Structures:
4G8G, 4G8I, 4G9D, 4G9F - PubMed Abstract:
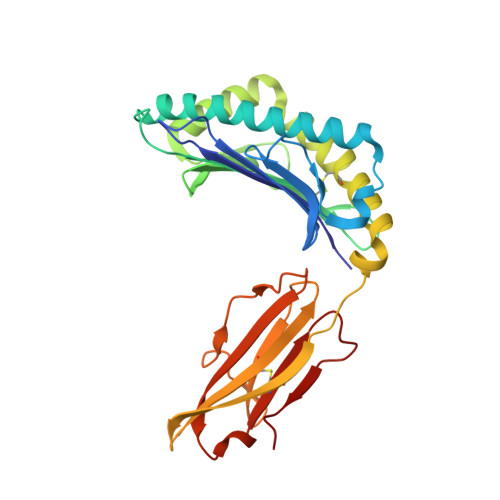
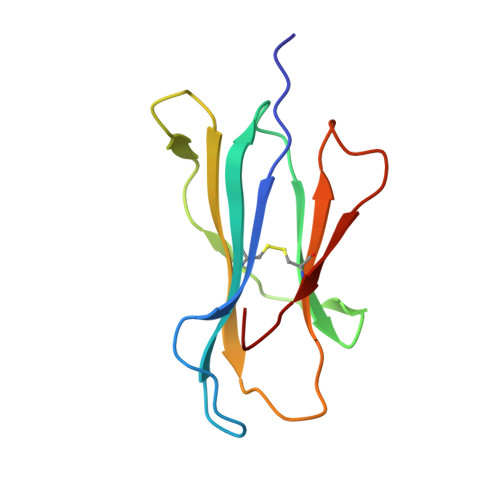
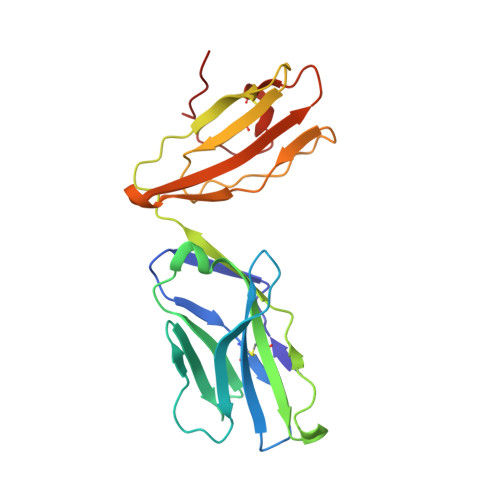
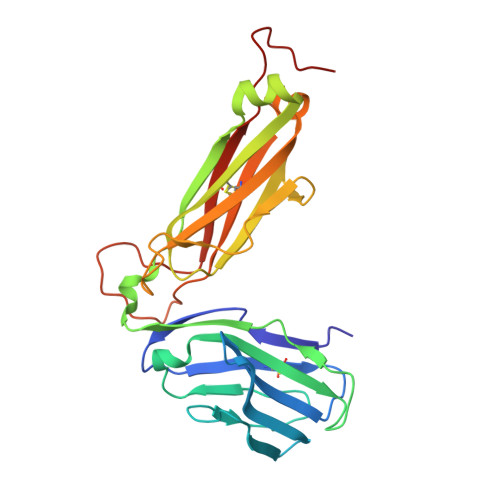
The capacity of the immune system to adapt to rapidly evolving viruses is a primary feature of effective immunity, yet its molecular basis is unclear. Here, we investigated protective HIV-1-specific CD8+ T cell responses directed against the immunodominant p24 Gag-derived epitope KK10 (KRWIILGLNK263-272) presented by human leukocyte antigen (HLA)-B∗2705. We found that cross-reactive CD8+ T cell clonotypes were mobilized to counter the rapid emergence of HIV-1 variants that can directly affect T cell receptor (TCR) recognition. These newly recruited clonotypes expressed TCRs that engaged wild-type and mutant KK10 antigens with similar affinities and almost identical docking modes, thereby accounting for their antiviral efficacy in HLA-B∗2705+ individuals. A protective CD8+ T cell repertoire therefore encompasses the capacity to control TCR-accessible mutations, ultimately driving the development of more complex viral escape variants that disrupt antigen presentation.
- Institute of Infection and Immunity, Cardiff University School of Medicine, Heath Park, Cardiff CF14 4XN, Wales, UK.
Organizational Affiliation: