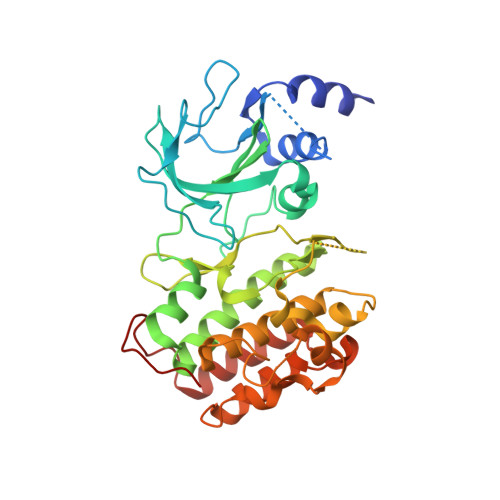
The crystal structure of the catalytic domain of the NF-kappaB inducing kinase reveals a narrow but flexible active site.
de Leon-Boenig, G., Bowman, K.K., Feng, J.A., Crawford, T., Everett, C., Franke, Y., Oh, A., Stanley, M., Staben, S.T., Starovasnik, M.A., Wallweber, H.J., Wu, J., Wu, L.C., Johnson, A.R., Hymowitz, S.G.(2012) Structure 20: 1704-1714
- PubMed: 22921830 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2012.07.013
- Primary Citation Related Structures:
4G3C, 4G3D, 4G3E, 4G3F, 4G3G - PubMed Abstract:
The NF-κB inducing kinase (NIK) regulates the non-canonical NF-κB pathway downstream of important clinical targets including BAFF, RANKL, and LTβ. Despite numerous genetic studies associating dysregulation of this pathway with autoimmune diseases and hematological cancers, detailed molecular characterization of this central signaling node has been lacking. We undertook a systematic cloning and expression effort to generate soluble, well-behaved proteins encompassing the kinase domains of human and murine NIK. Structures of the apo NIK kinase domain from both species reveal an active-like conformation in the absence of phosphorylation. ATP consumption and peptide phosphorylation assays confirm that phosphorylation of NIK does not increase enzymatic activity. Structures of murine NIK bound to inhibitors possessing two different chemotypes reveal conformational flexibility in the gatekeeper residue controlling access to a hydrophobic pocket. Finally, a single amino acid difference affects the ability of some inhibitors to bind murine and human NIK with the same affinity.
- Department of Structural Biology, Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, USA.
Organizational Affiliation: