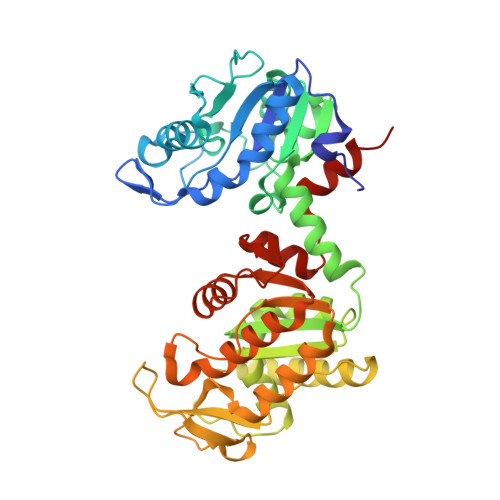
An X-ray Structure of a Putative Phosphogylcerate Kinase with Bound ADP from Francisella tularensis subsp. tularensis SCHU S4
Brunzelle, J.S., Wawrzak, Z., Skarina, T., Anderson, W.F., Savchenko, A., Center for Structural Genomics of Infectious DiseasesTo be published.