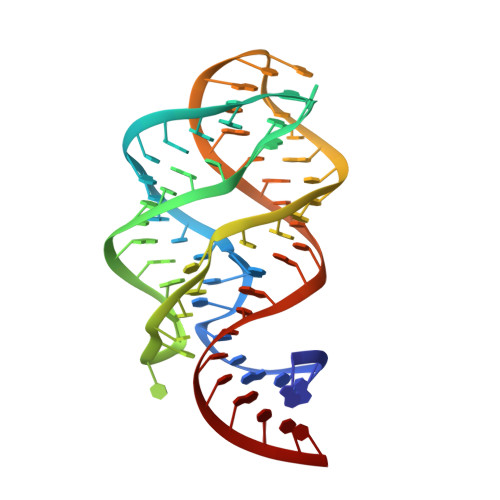
Structure of a natural guanine-responsive riboswitch complexed with the metabolite hypoxanthine.
Batey, R.T., Gilbert, S.D., Montange, R.K.(2004) Nature 432: 411-415
- PubMed: 15549109 Search on PubMed
- DOI: https://doi.org/10.1038/nature03037
- Primary Citation Related Structures:
4FE5 - PubMed Abstract:
Riboswitches are genetic regulatory elements found in the 5' untranslated region of messenger RNA that act in the absence of protein cofactors. They are broadly distributed across bacteria and account for the regulation of more than 2% of all genes in Bacillus subtilis, underscoring their importance in the control of cellular metabolism. The 5' untranslated region of many mRNAs of genes involved in purine metabolism and transport contain a guanine-responsive riboswitch that directly binds guanine, hypoxanthine or xanthine to terminate transcription. Here we report the crystal structure at 1.95 A resolution of the purine-binding domain of the guanine riboswitch from the xpt-pbuX operon of B. subtilis bound to hypoxanthine, a prevalent metabolite in the bacterial purine salvage pathway. This structure reveals a complex RNA fold involving several phylogenetically conserved nucleotides that create a binding pocket that almost completely envelops the ligand. Hypoxanthine functions to stabilize this structure and to promote the formation of a downstream transcriptional terminator element, thereby providing a mechanism for directly repressing gene expression in response to an increase in intracellular concentrations of metabolite.
- Department of Chemistry and Biochemistry, 215 UCB, University of Colorado, Boulder, Colorado 80309, USA. robert.batey@colorado.edu
Organizational Affiliation: