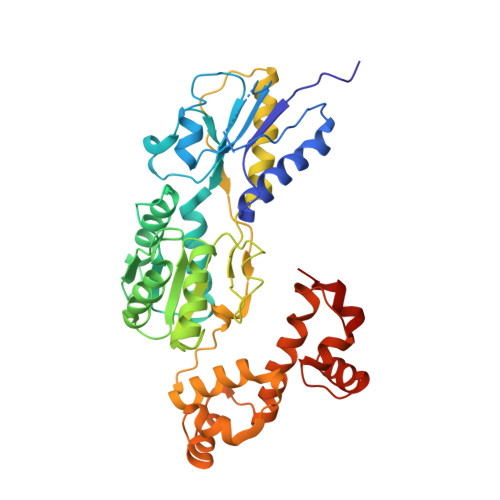
Structures of the Escherichia coli transcription activator and regulator of diauxie, XylR: an AraC DNA-binding family member with a LacI/GalR ligand-binding domain.
Ni, L., Tonthat, N.K., Chinnam, N., Schumacher, M.A.(2013) Nucleic Acids Res 41: 1998-2008
- PubMed: 23241389 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gks1207
- Primary Citation Related Structures:
4FE4, 4FE7 - PubMed Abstract:
Escherichia coli can rapidly switch to the metabolism of l-arabinose and d-xylose in the absence of its preferred carbon source, glucose, in a process called carbon catabolite repression. Transcription of the genes required for l-arabinose and d-xylose consumption is regulated by the sugar-responsive transcription factors, AraC and XylR. E. coli represents a promising candidate for biofuel production through the metabolism of hemicellulose, which is composed of d-xylose and l-arabinose. Understanding the l-arabinose/d-xylose regulatory network is key for such biocatalyst development. Unlike AraC, which is a well-studied protein, little is known about XylR. To gain insight into XylR function, we performed biochemical and structural studies. XylR contains a C-terminal AraC-like domain. However, its N-terminal d-xylose-binding domain contains a periplasmic-binding protein (PBP) fold with structural homology to LacI/GalR transcription regulators. Like LacI/GalR proteins, the XylR PBP domain mediates dimerization. However, unlike LacI/GalR proteins, which dimerize in a parallel, side-to-side manner, XylR PBP dimers are antiparallel. Strikingly, d-xylose binding to this domain results in a helix to strand transition at the dimer interface that reorients both DNA-binding domains, allowing them to bind and loop distant operator sites. Thus, the combined data reveal the ligand-induced activation mechanism of a new family of DNA-binding proteins.
- Department of Biochemistry and Molecular Biology, University of Texas, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
Organizational Affiliation: