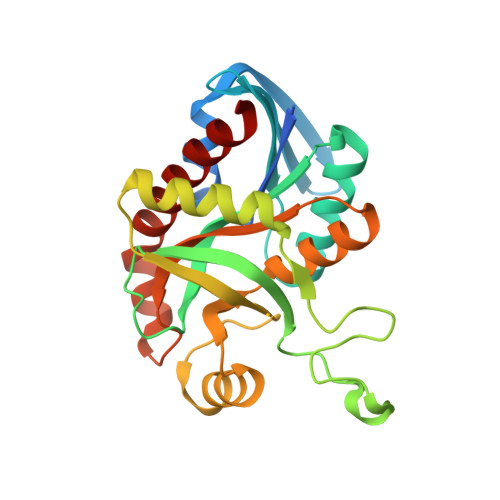
Crystal structure of 5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase from Salmonella enterica with diEtglycol-thio-DADMe-Immucillin-A
Haapalainen, A.M., Thomas, K., Tyler, P.C., Evans, G.B., Almo, S.C., Schramm, V.L.(2013) Structure 21: 963-974
- PubMed: 23685211 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2013.04.009
- Primary Citation Related Structures:
4F1W, 4F2P, 4F2W, 4F3C, 4F3K - PubMed Abstract:
Accumulation of 5'-methylthioadenosine (MTA) and S-adenosylhomocysteine (SAH) in bacteria disrupts the S-adenosylmethionine pool to alter biological methylations, synthesis of polyamines, and production of quorum-sensing molecules. Bacterial metabolism of MTA and SAH depends on MTA/SAH nucleosidase (MTAN), an enzyme not present in humans and a target for quorum sensing because MTAN activity is essential for synthesis of autoinducer-2 molecules. Crystals of Salmonella enterica MTAN with product and transition state analogs of MTA and SAH explain the structural contacts causing pM binding affinity for the inhibitor and reveal a "water-wire" channel for the catalytic nucleophile. The crystal structure shows an extension of the binding pocket filled with polyethylene glycol. We exploited this discovery by the design and synthesis of tailored modifications of the currently existing transition state analogs to fill this site. This site was not anticipated in MTAN structures. Tailored inhibitors with dissociation constants of 5 to 15 pM are characterized.
- Department of Biochemistry, Albert Einstein College of Medicine of Yeshiva University, 1300 Morris Park Avenue, Bronx, NY 10461, USA.
Organizational Affiliation: