A Selective and Brain Penetrant p38 alpha MAPK Inhibitor Candidate for Neurologic and Neuropsychiatric Disorders That Attenuates Neuroinflammation and Cognitive Dysfunction.
Roy, S.M., Minasov, G., Arancio, O., Chico, L.W., Van Eldik, L.J., Anderson, W.F., Pelletier, J.C., Watterson, D.M.(2019) J Med Chem 62: 5298-5311
- PubMed: 30978288 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.9b00058
- Primary Citation Related Structures:
4EWQ, 4ZTH - PubMed Abstract:
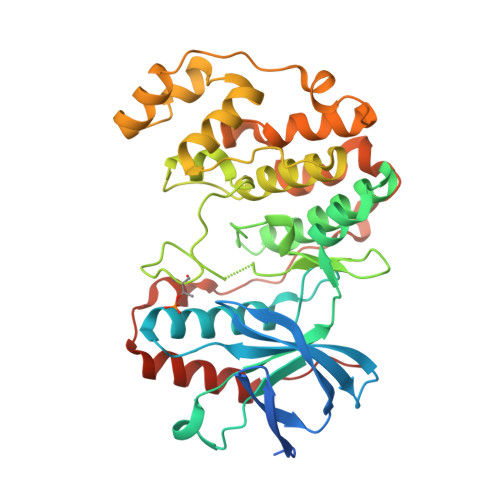
The p38αMAPK is a serine/threonine protein kinase and a key node in the intracellular signaling networks that transduce and amplify stress signals into physiological changes. A preponderance of preclinical data and clinical observations established p38αMAPK as a brain drug discovery target involved in neuroinflammatory responses and synaptic dysfunction in multiple degenerative and neuropsychiatric brain disorders. We summarize the discovery of highly selective, brain-penetrant, small molecule p38αMAPK inhibitors that are efficacious in diverse animal models of neurologic disorders. A crystallography and pharmacoinformatic approach to fragment expansion enabled the discovery of an efficacious hit. The addition of secondary pharmacology screens to refinement delivered lead compounds with improved selectivity, appropriate pharmacodynamics, and efficacy. Safety considerations and additional secondary pharmacology screens drove optimization that delivered the drug candidate MW01-18-150SRM (MW150), currently in early stage clinical trials.
- Northwestern University , 320 East Superior Street , Chicago , Illinois 60611 , United States.
Organizational Affiliation: