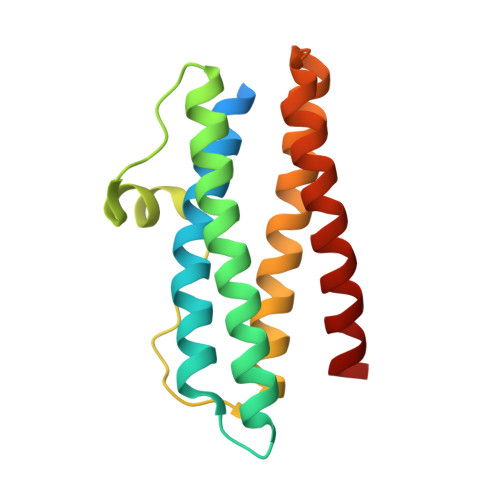
Crystal structure of Helicobacter pylori neutrophil-activating protein with a di-nuclear ferroxidase center in a zinc or cadmium-bound form
Yokoyama, H., Tsuruta, O., Akao, N., Fujii, S.(2012) Biochem Biophys Res Commun 422: 745-750
- PubMed: 22618234 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2012.05.073
- PubMed Abstract:
Helicobacter pylori neutrophil-activating protein (HP-NAP) is a Dps-like iron storage protein forming a dodecameric shell, and promotes adhesion of neutrophils to endothelial cells. The crystal structure of HP-NAP in a Zn(2+)- or Cd(2+)-bound form reveals the binding of two zinc or two cadmium ions and their bridged water molecule at the ferroxidase center (FOC). The two zinc ions are coordinated in a tetrahedral manner to the conserved residues among HP-NAP and Dps proteins. The two cadmium ions are coordinated in a trigonal-bipyramidal and distorted octahedral manner. In both structures, the second ion is more weakly coordinated than the first. Another zinc ion is found inside of the negatively-charged threefold-related pore, which is suitable for metal ions to pass through.
- School of Pharmaceutical Sciences, University of Shizuoka, 52-1 Yada, Shizuoka 422-8526, Japan. h-yokoya@u-shizuoka-ken.ac.jp
Organizational Affiliation: