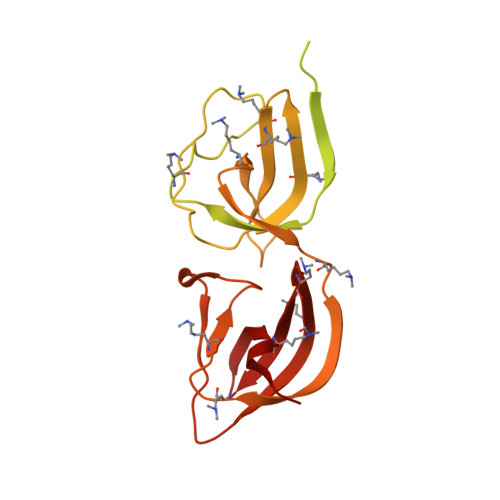
Ligand-binding properties and conformational dynamics of autolysin repeat domains in staphylococcal cell wall recognition.
Zoll, S., Schlag, M., Shkumatov, A.V., Rautenberg, M., Svergun, D.I., Gotz, F., Stehle, T.(2012) J Bacteriol 194: 3789-3802
- PubMed: 22609916 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00331-12
- Primary Citation Related Structures:
4EPC - PubMed Abstract:
The bifunctional major autolysin Atl plays a key role in staphylococcal cell separation. Processing of Atl yields catalytically active amidase (AM) and glucosaminidase (GL) domains that are each fused to repeating units. The two repeats of AM (R1 and R2) target the enzyme to the septum, where it cleaves murein between dividing cells. We have determined the crystal structure of R2, which reveals that each repeat folds into two half-open β-barrel subunits. We further demonstrate that lipoteichoic acid serves as a receptor for the repeats and that this interaction depends on conserved surfaces in each subunit. Small-angle X-ray scattering of the mature amidase reveals the presence of flexible linkers separating the AM, R1, and R2 units. Different levels of flexibility for each linker provide mechanistic insights into the conformational dynamics of the full-length protein and the roles of its components in cell wall association and catalysis. Our analysis supports a model in which the repeats direct the catalytic AM domain to the septum, where it can optimally perform the final step of cell division.
- Interfaculty Institute of Biochemistry, University of Tuebingen, Tuebingen, Germany.
Organizational Affiliation: