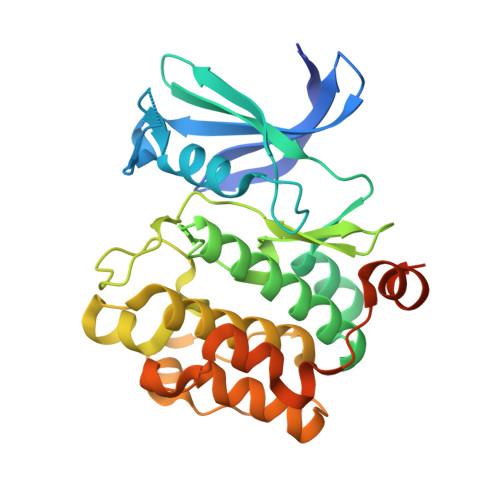
Flexibility of the P-loop of Pim-1 kinase: observation of a novel conformation induced by interaction with an inhibitor
Parker, L.J., Watanabe, H., Tsuganezawa, K., Tomabechi, Y., Handa, N., Shirouzu, M., Yuki, H., Honma, T., Ogawa, N., Nagano, T., Yokoyama, S., Tanaka, A.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 860-866
- PubMed: 22869110 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309112027108
- Primary Citation Related Structures:
3UMX, 4ENX, 4ENY - PubMed Abstract:
The serine/threonine kinase Pim-1 is emerging as a promising target for cancer therapeutics. Much attention has recently been focused on identifying potential Pim-1 inhibitor candidates for the treatment of haematopoietic malignancies. The outcome of a rational drug-design project has recently been reported [Nakano et al. (2012), J. Med. Chem. 55, 5151-5156]. The report described the process of optimization of the structure-activity relationship and detailed from a medicinal chemistry perspective the development of a low-potency and nonselective compound initially identified from in silico screening into a potent, selective and metabolically stable Pim-1 inhibitor. Here, the structures of the initial in silico hits are reported and the noteworthy features of the Pim-1 complex structures are described. A particular focus was placed on the rearrangement of the glycine-rich P-loop region that was observed for one of the initial compounds, (Z)-7-(azepan-1-ylmethyl)-2-[(1H-indol-3-yl)methylidene]-6-hydroxy-1-benzofuran-3(2H)-one (compound 1), and was also found in all further derivatives. This novel P-loop conformation, which appears to be stabilized by an additional interaction with the β3 strand located above the binding site, is not usually observed in Pim-1 structures.
- RIKEN Systems and Structural Biology Center, 1-7-22 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: