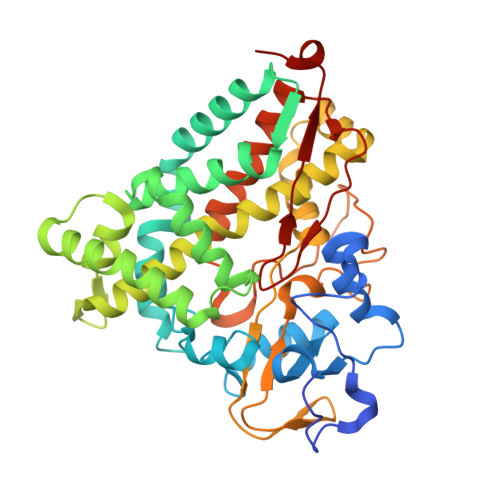
Double electron-electron resonance shows cytochrome P450cam undergoes a conformational change in solution upon binding substrate.
Stoll, S., Lee, Y.T., Zhang, M., Wilson, R.F., Britt, R.D., Goodin, D.B.(2012) Proc Natl Acad Sci U S A 109: 12888-12893
- PubMed: 22826259 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1207123109
- Primary Citation Related Structures:
4EK1 - PubMed Abstract:
Although cytochrome P450cam from Pseudomonas putida, the archetype for all heme monooxygenases, has long been known to have a closed active site, recent reports show that the enzyme can also be crystallized in at least two clusters of open conformations. This suggests that the enzyme may undergo significant conformational changes during substrate binding and catalytic turnover. However, these conformations were observed in the crystalline state, and information is needed about the conformations that are populated in solution. In this study, double electron-electron resonance experiments were performed to observe substrate-induced changes in distance as measured by the dipolar coupling between spin labels introduced onto the surface of the enzyme on opposite sides of the substrate access channel. The double electron-electron resonance data show a decrease of 0.8 nm in the distance between spin labels placed at S48C and S190C upon binding the substrate camphor. A rotamer distribution model based on the crystal structures adequately describes the observed distance distributions. These results demonstrate conclusively that, in the physiologically relevant solution state, the substrate-free enzyme exists in the open P450cam-O conformation and that camphor binding results in conversion to the closed P450cam-C form. This approach should be useful for investigating many other P450s, including mammalian forms, in which the role of conformational change is of central importance but not well understood.
- Department of Chemistry, University of California, One Shields Avenue, Davis, CA 95616, USA.
Organizational Affiliation: